
Python Plotting for Exploratory Data Analysis
The simple graph has brought more information to the data analyst's mind than any other device.
Contents
- Introduction
- Examples
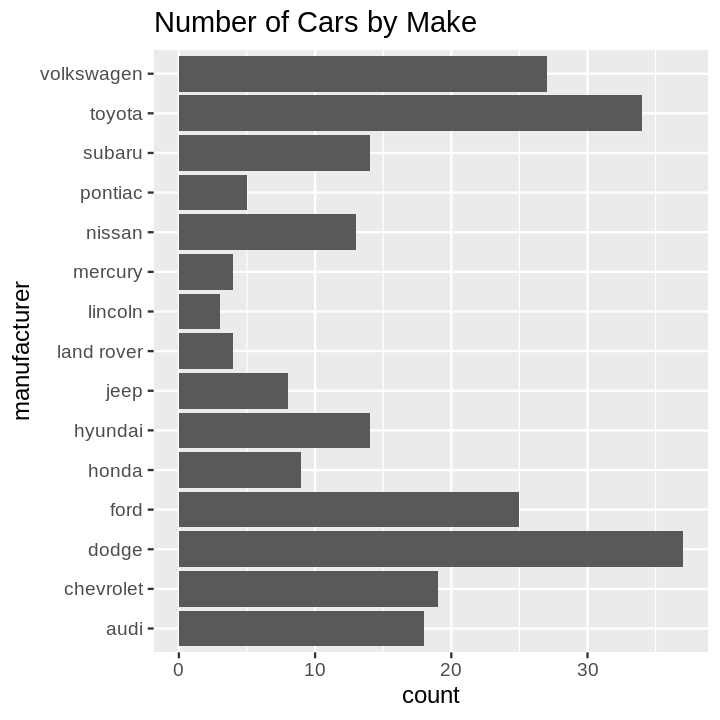
- Bar Chart
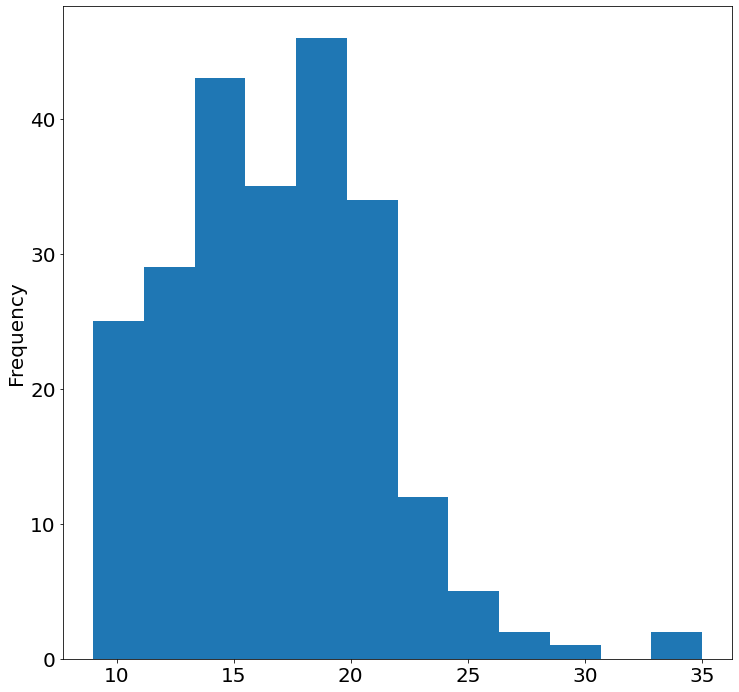
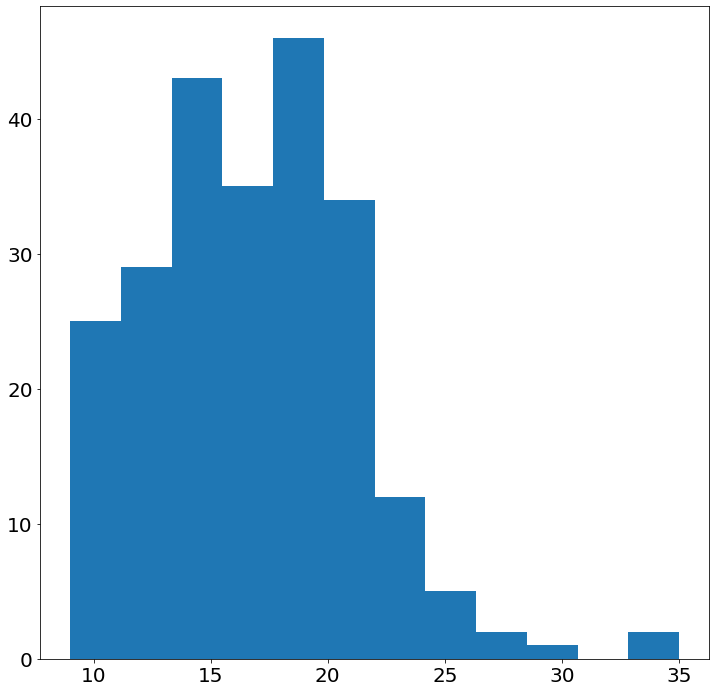
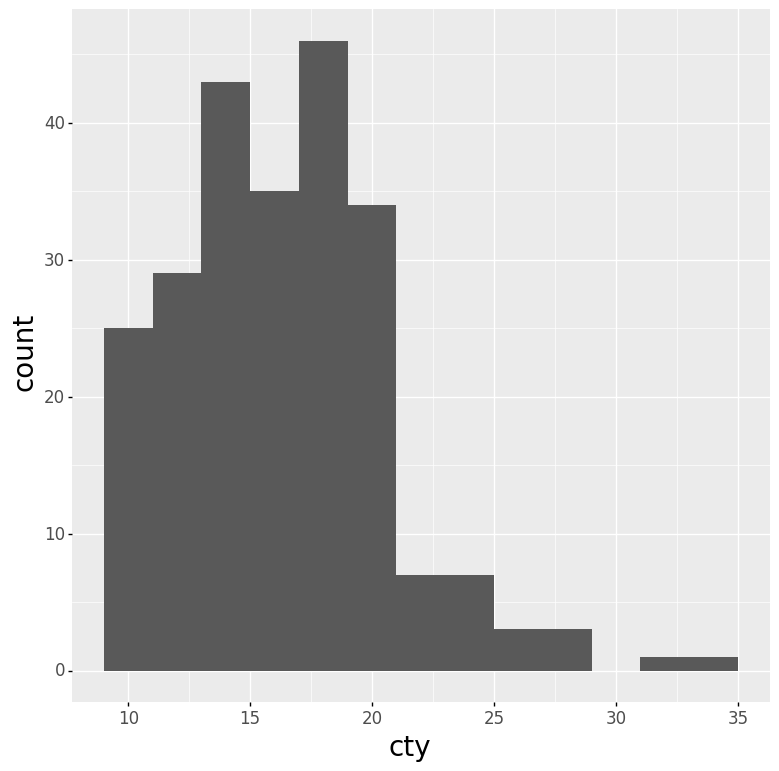
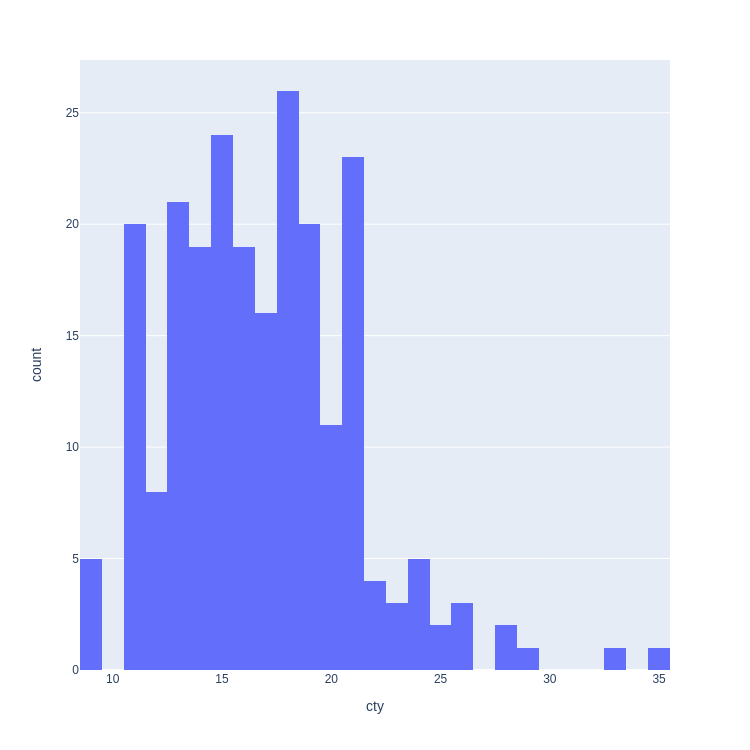
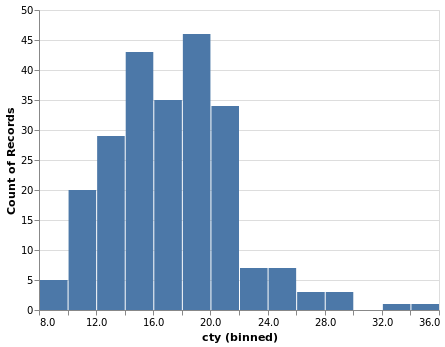
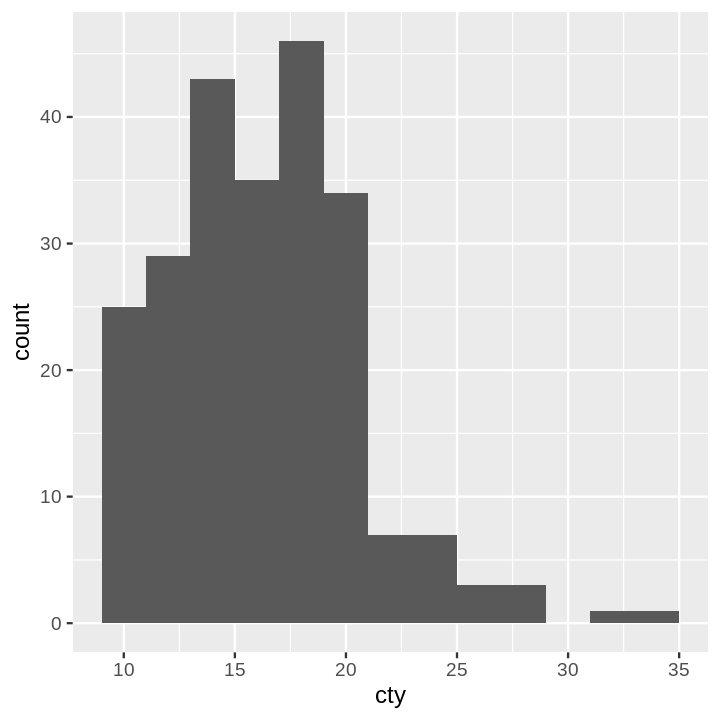
- Histogram
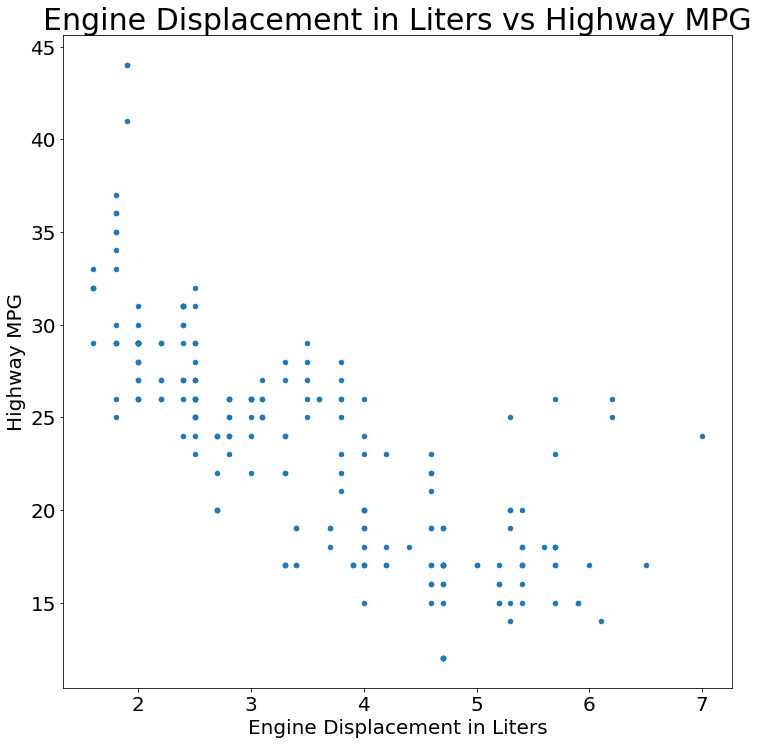
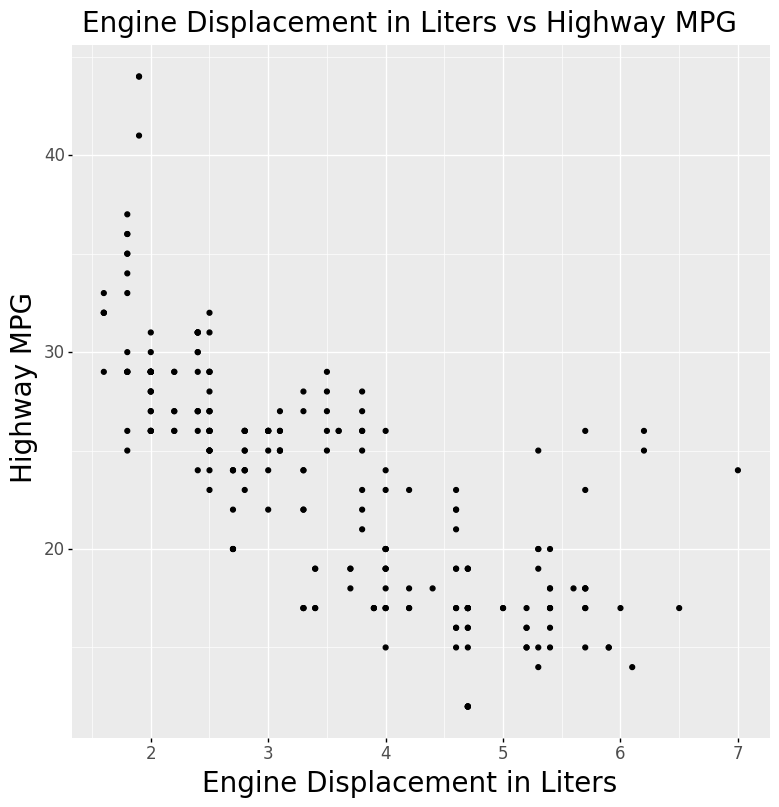
- Scatter Plot
- Time Series
- Scatter Plot with Faceted with Color
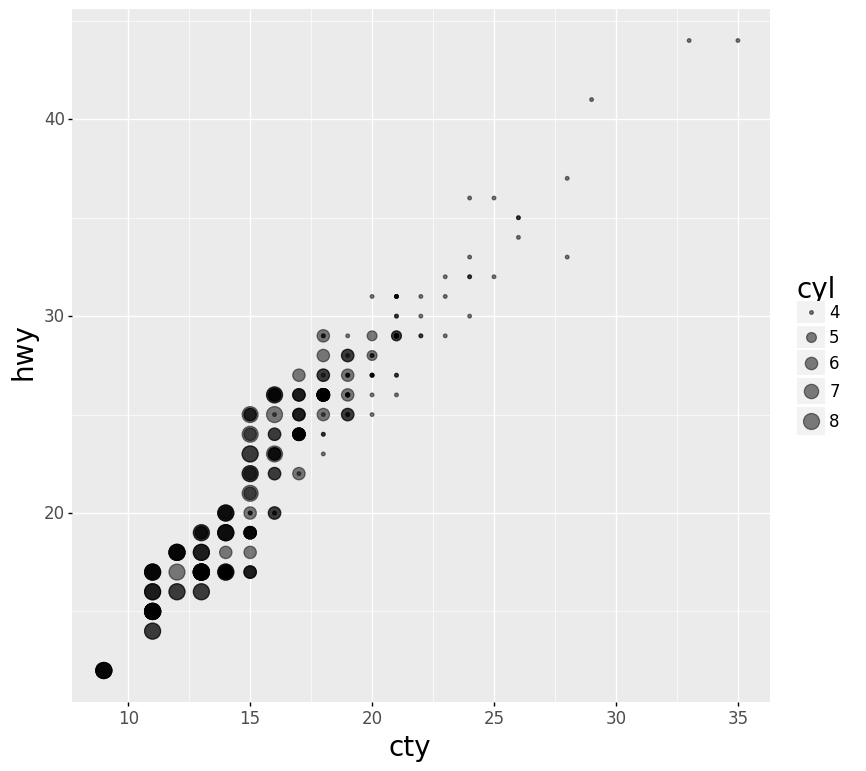
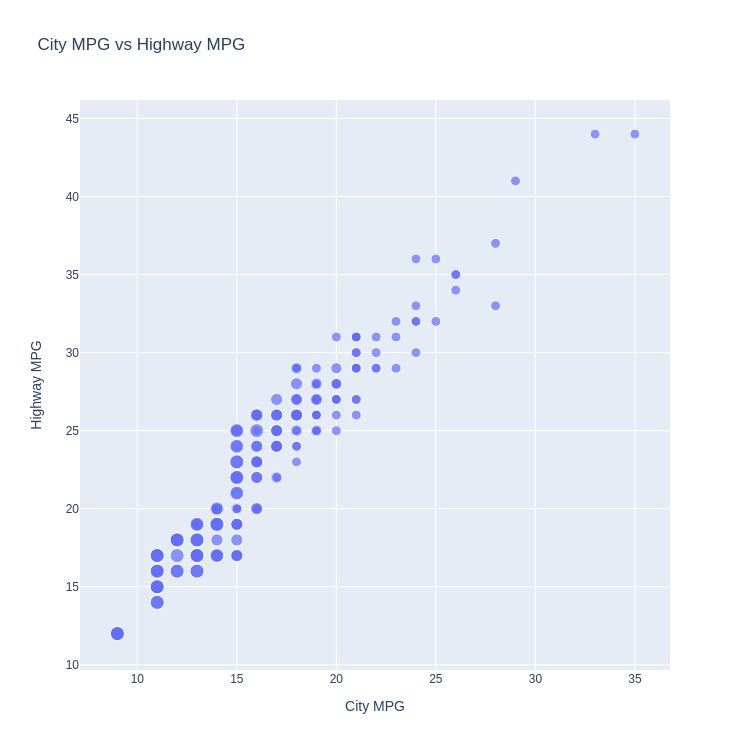
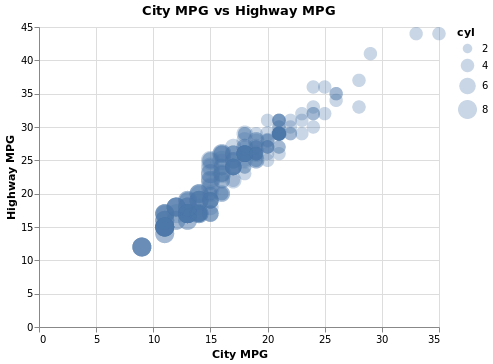
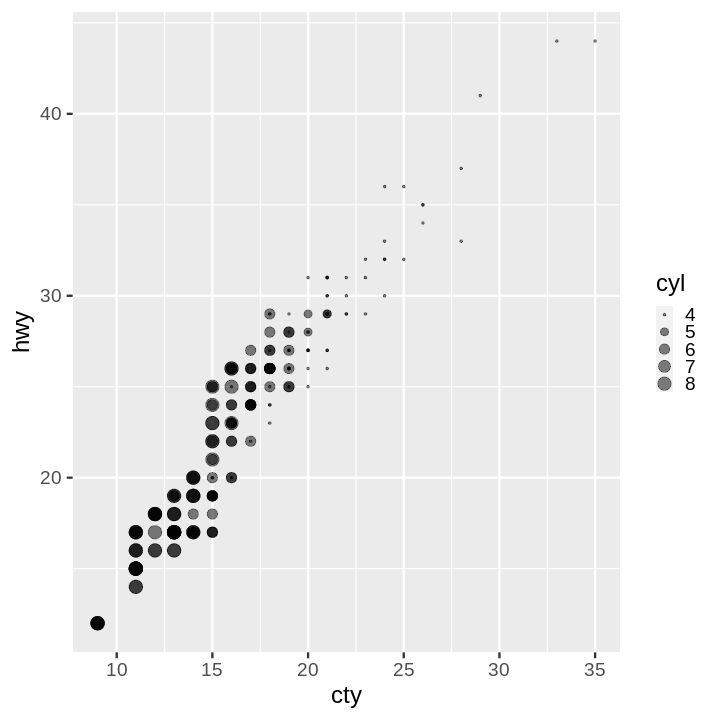
- Scatter Plot with Points Sized by Continuous Value
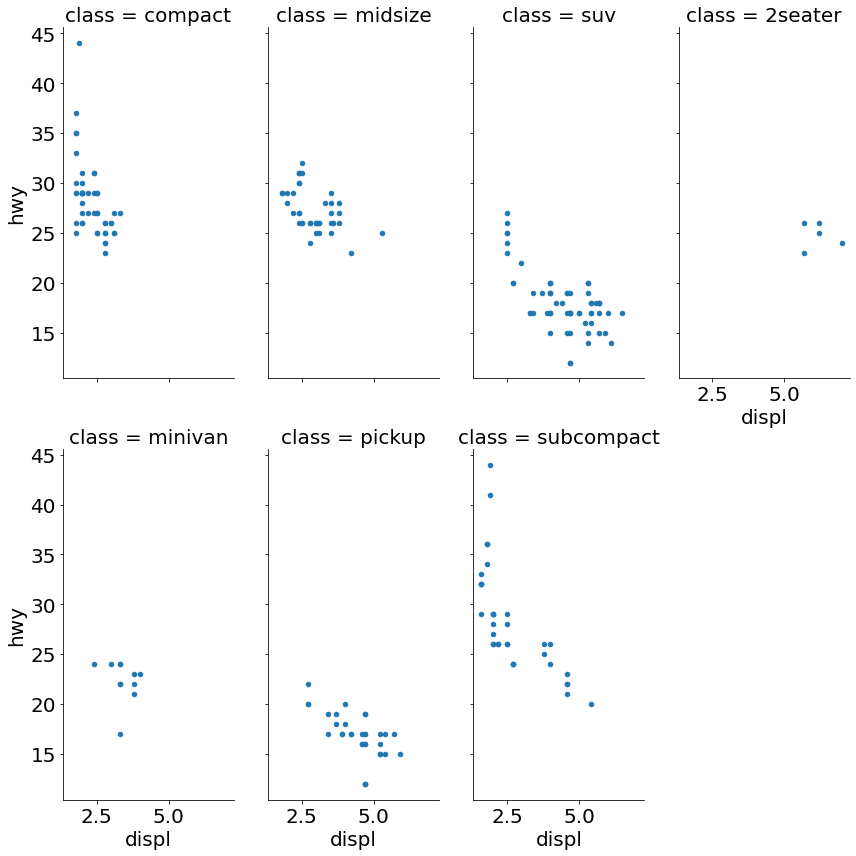
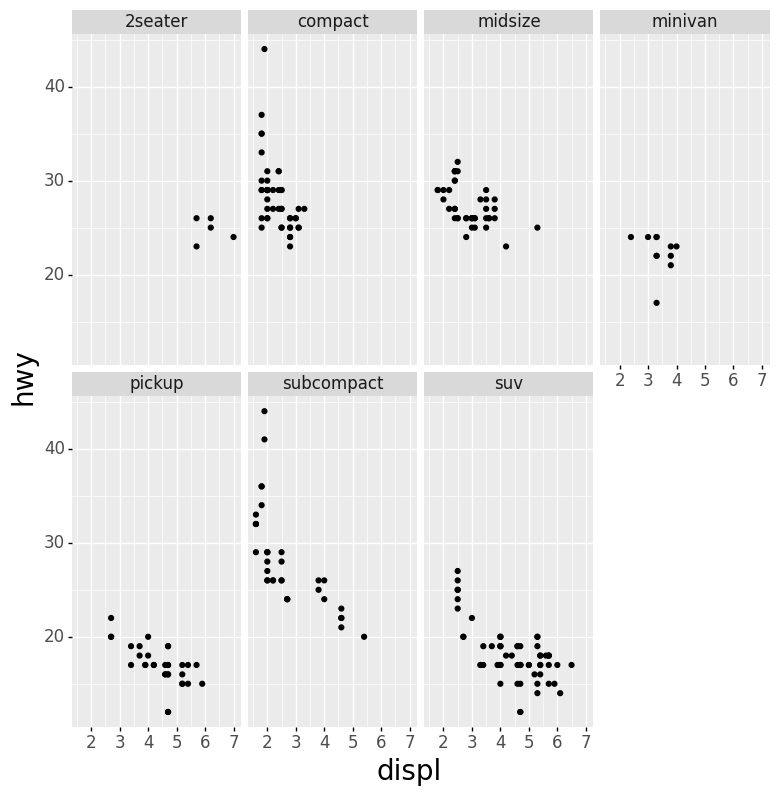
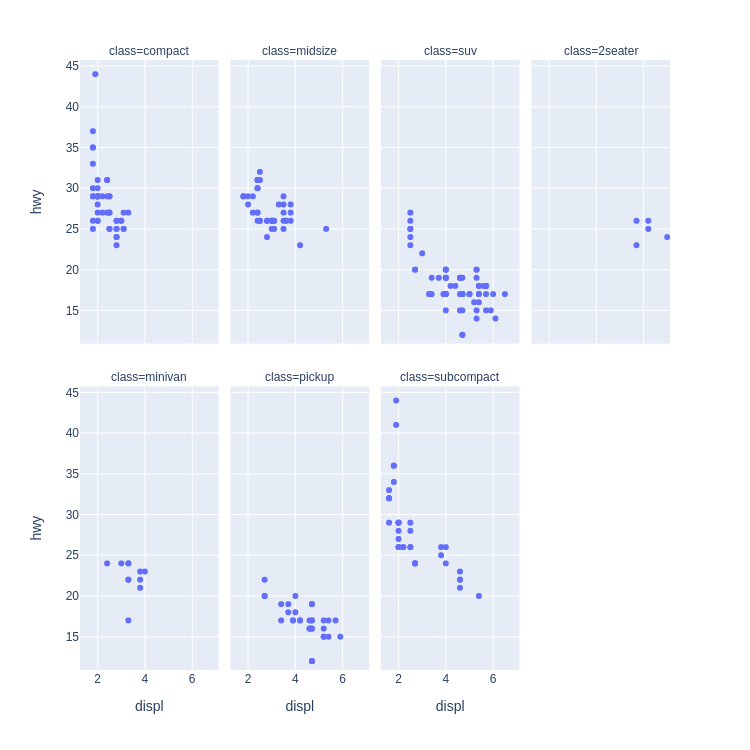
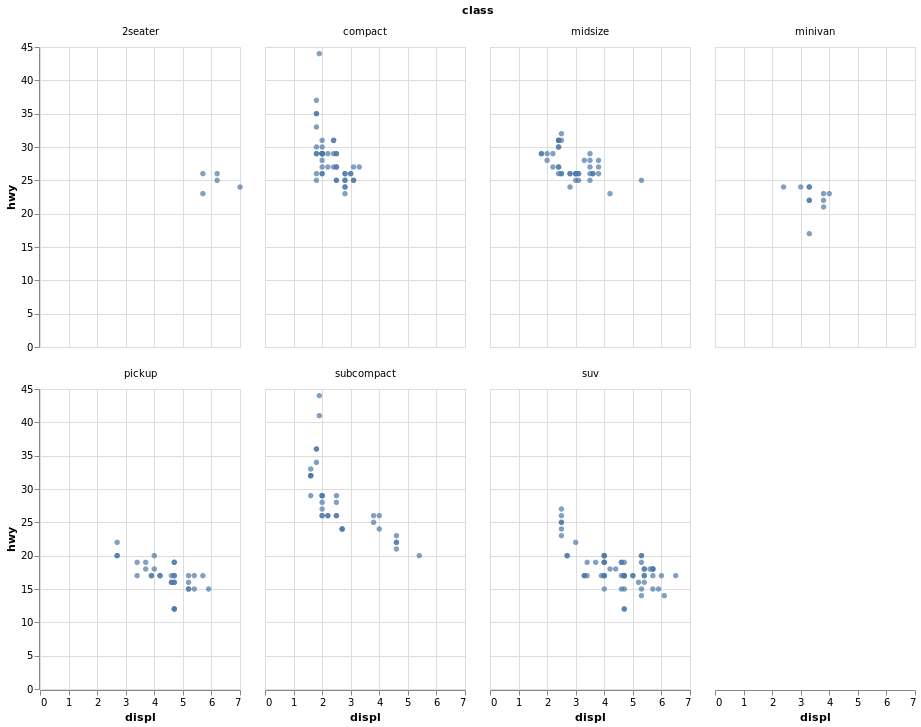
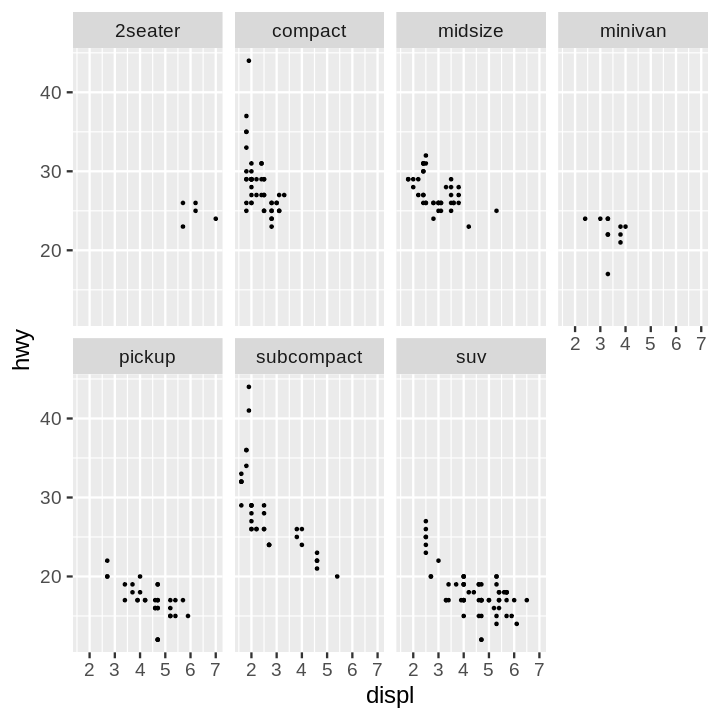
- Scatter Plot Faceted on One Variable
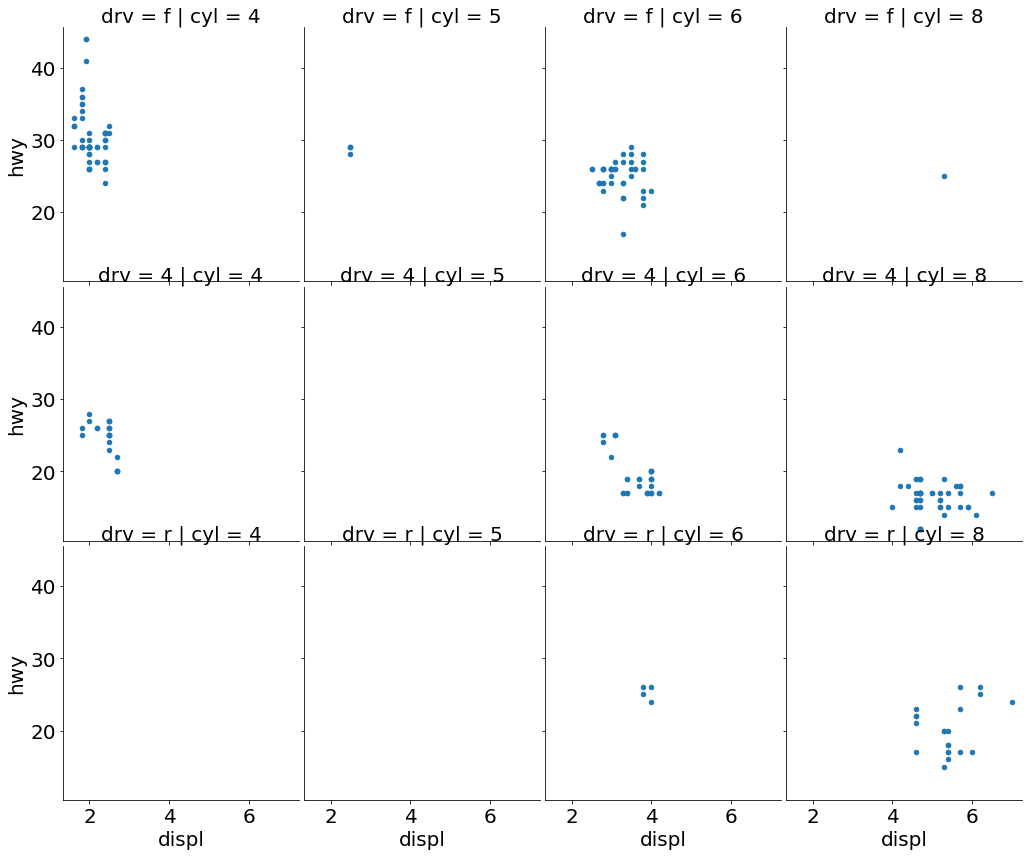
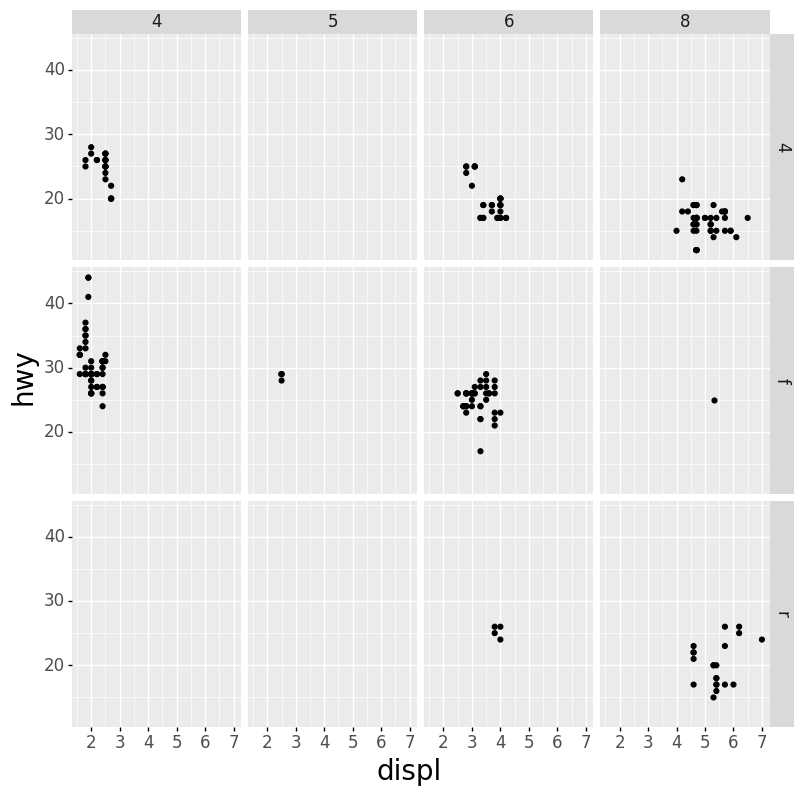
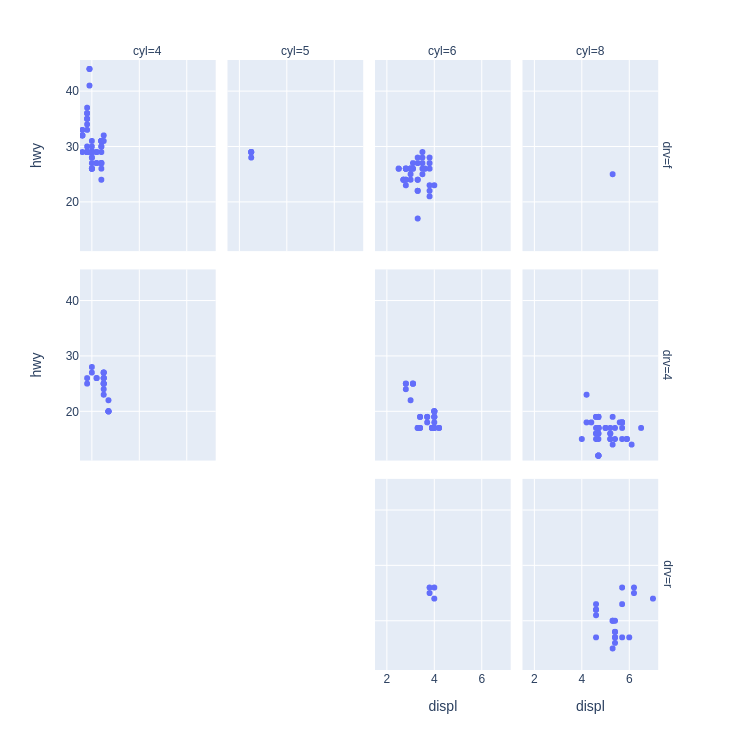
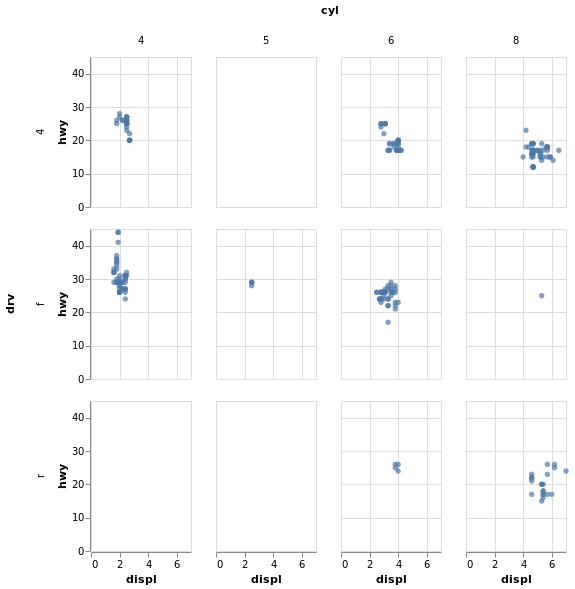
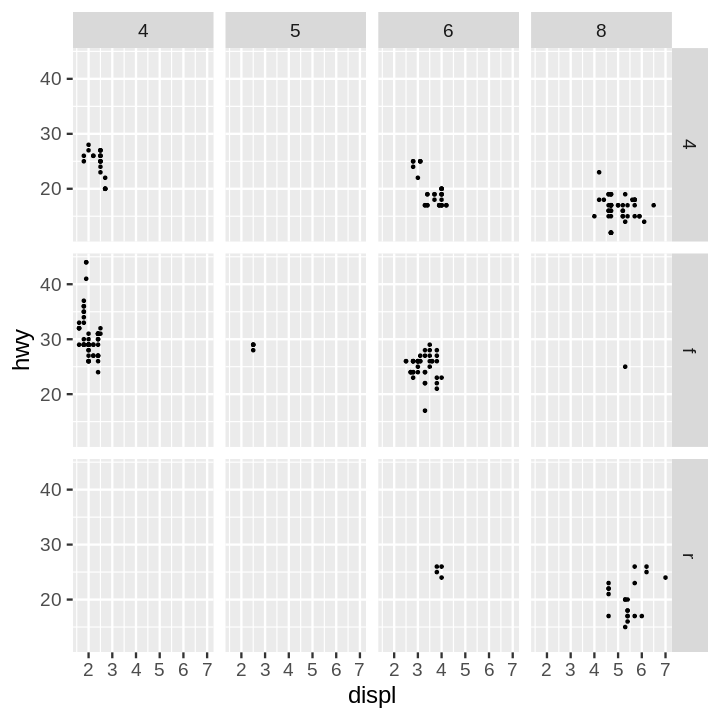
- Scatter Plot Faceted on Two Variables
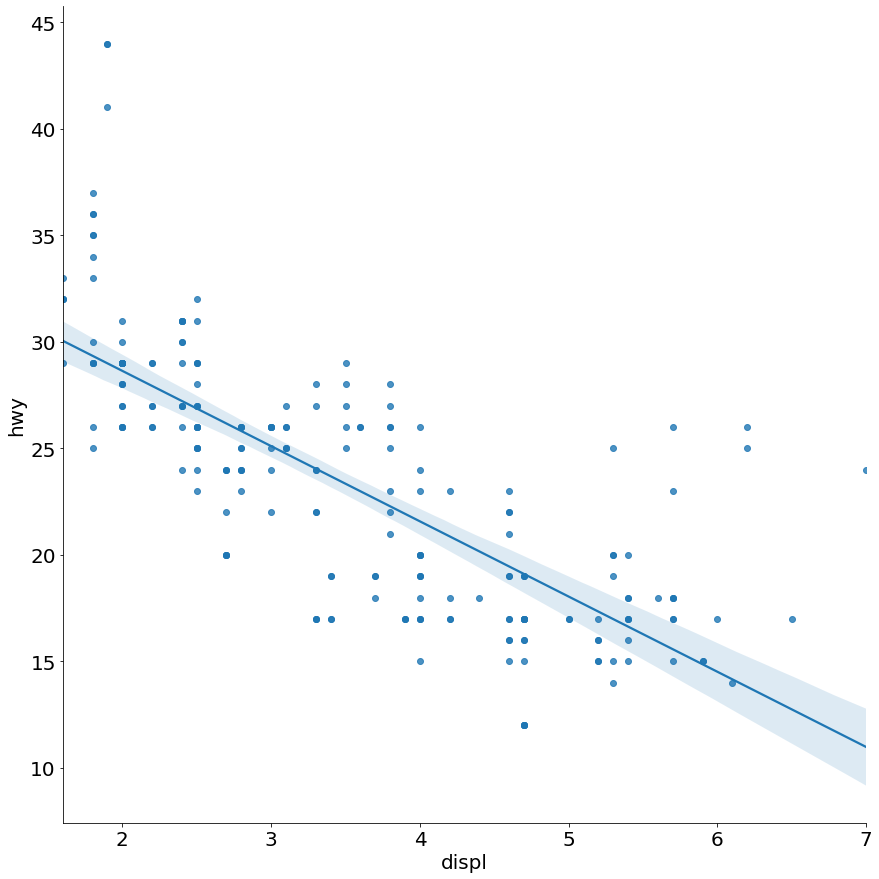
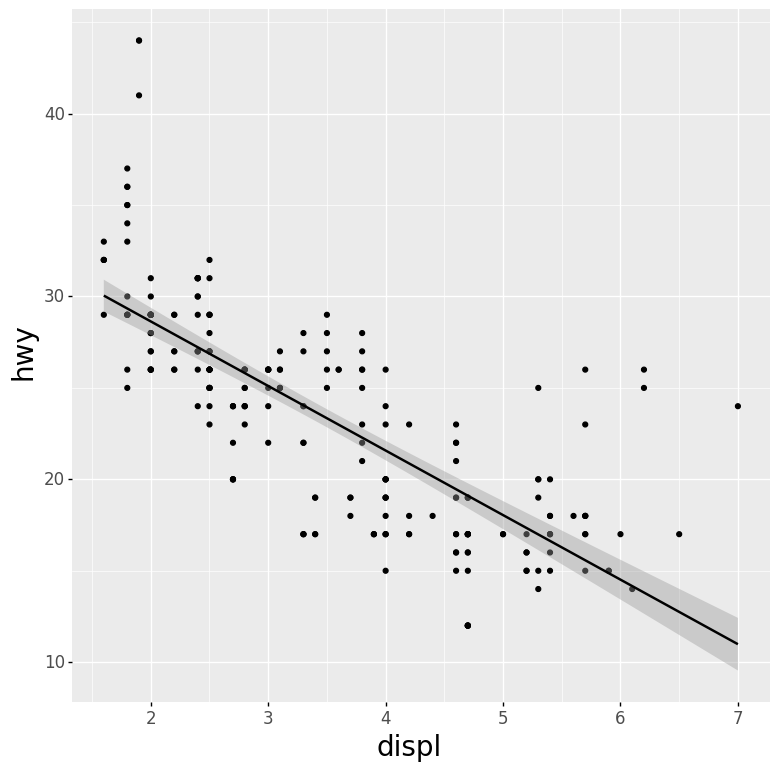
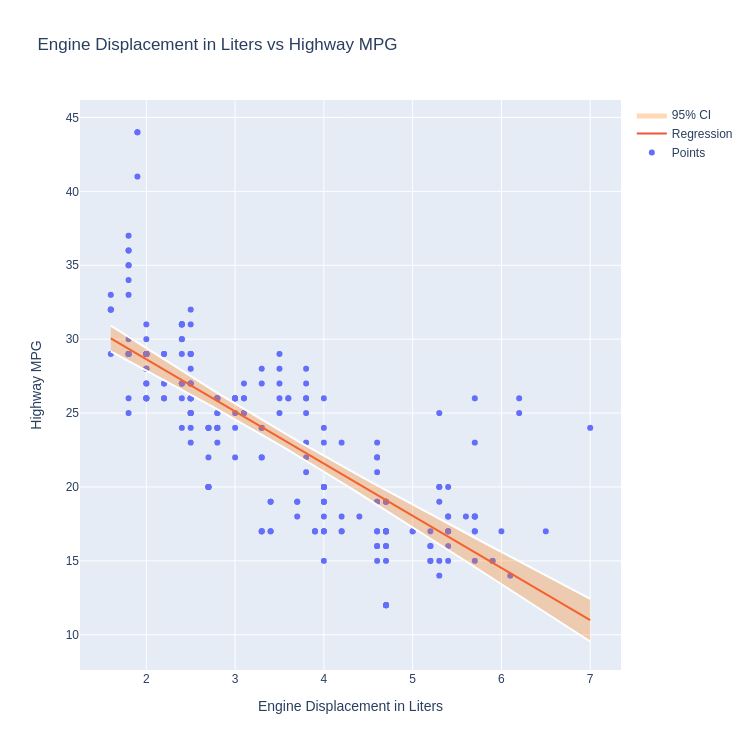
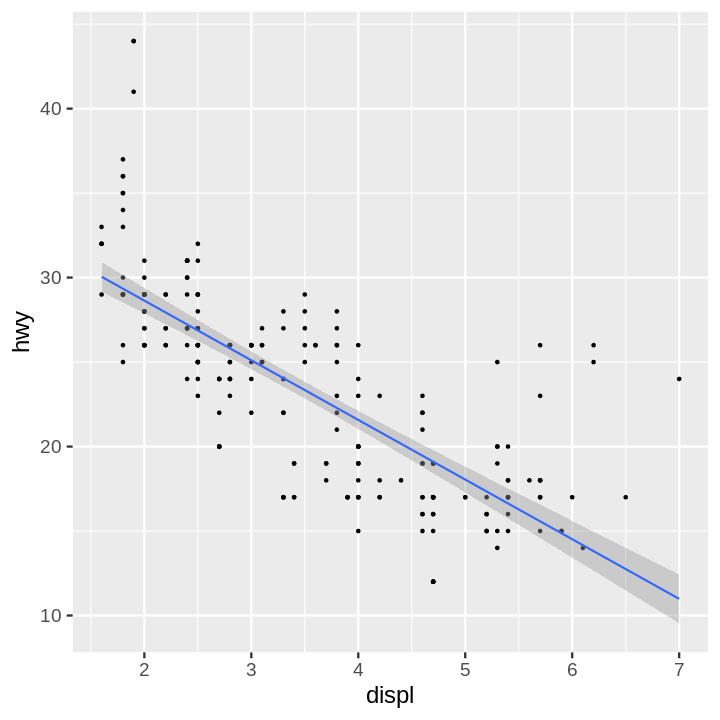
- Scatter Plot and Regression Line with 95% Confidence Interval Layered
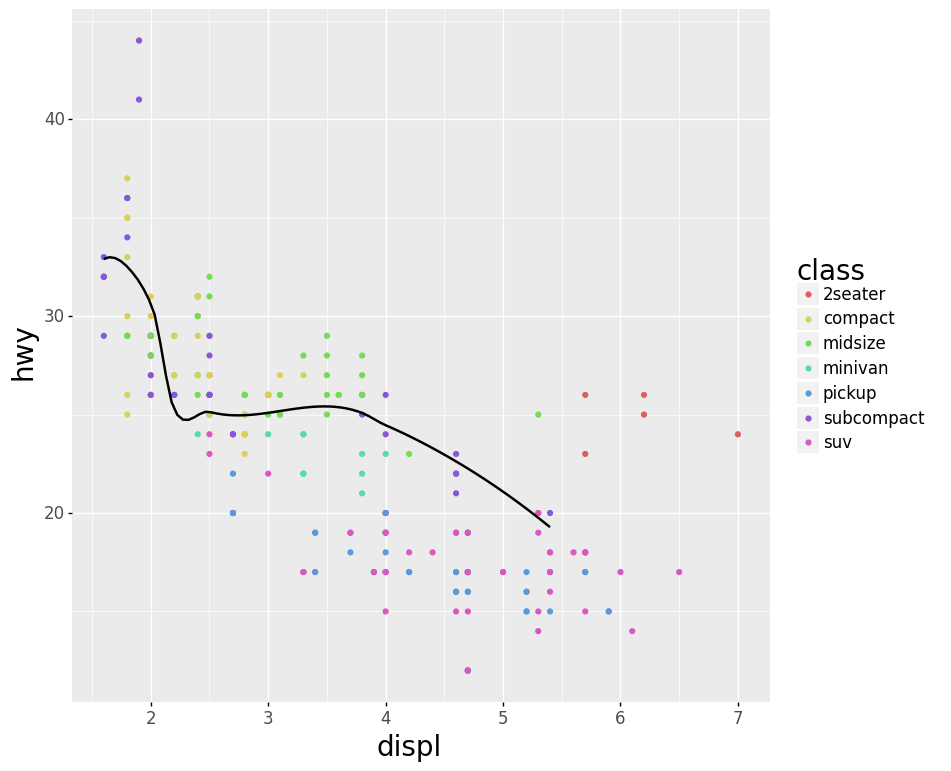
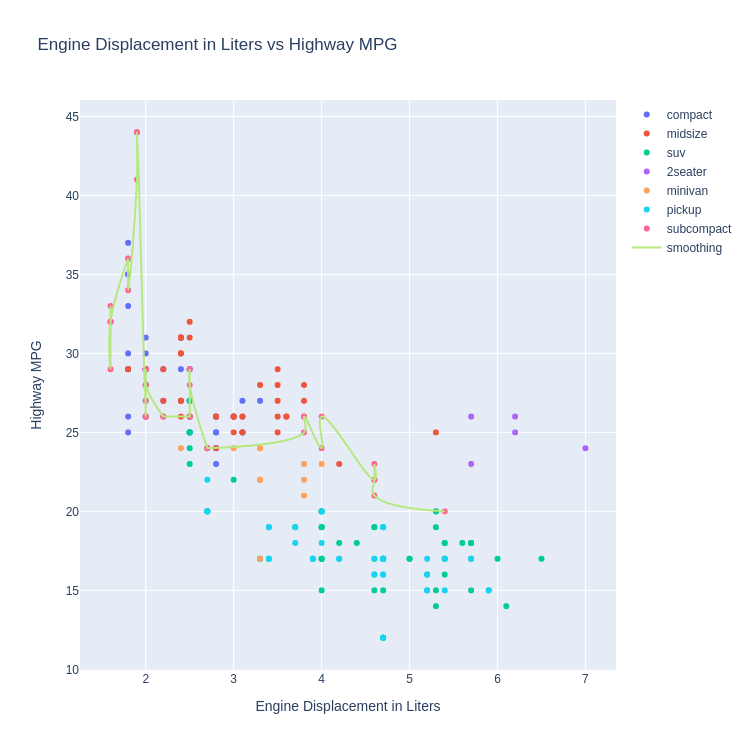
- Smoothed Line Plot and Scatter Plot Layered
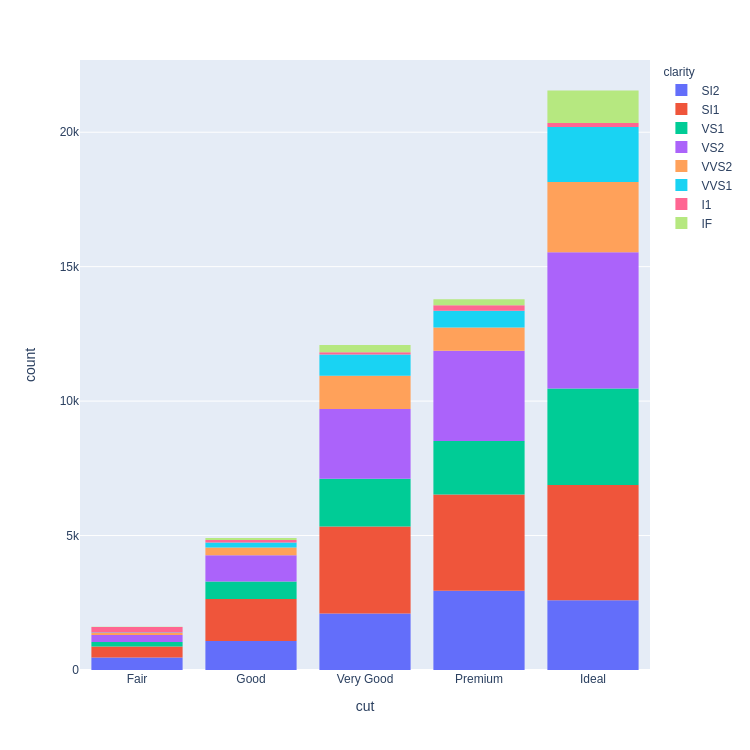
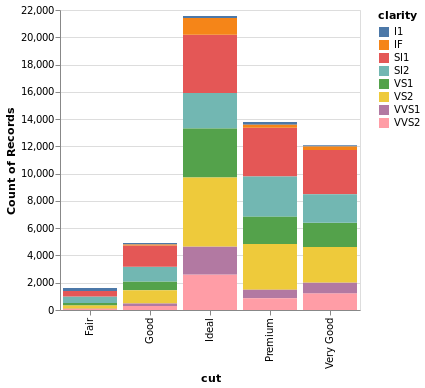
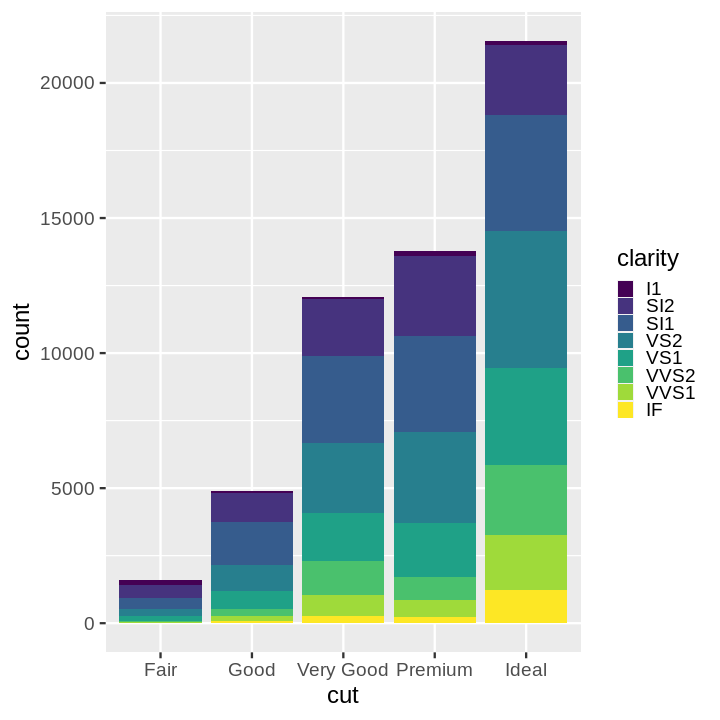
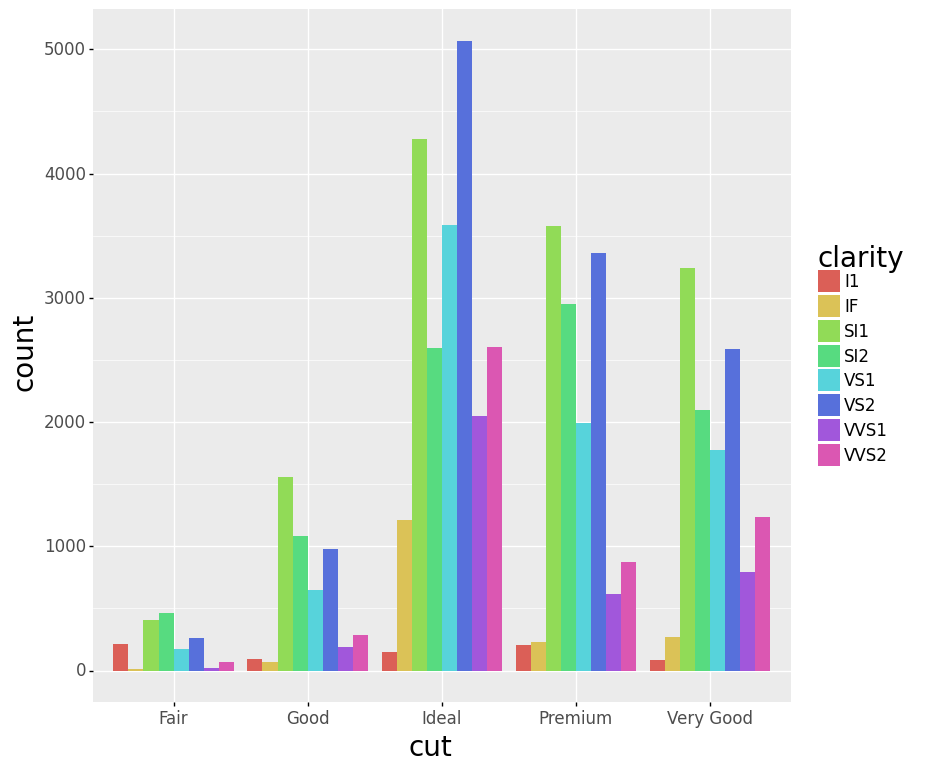
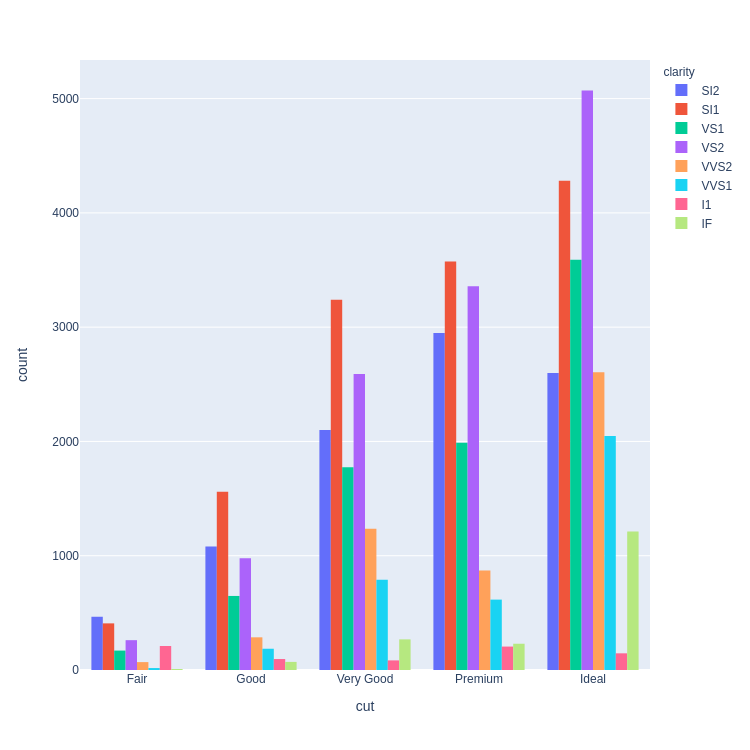
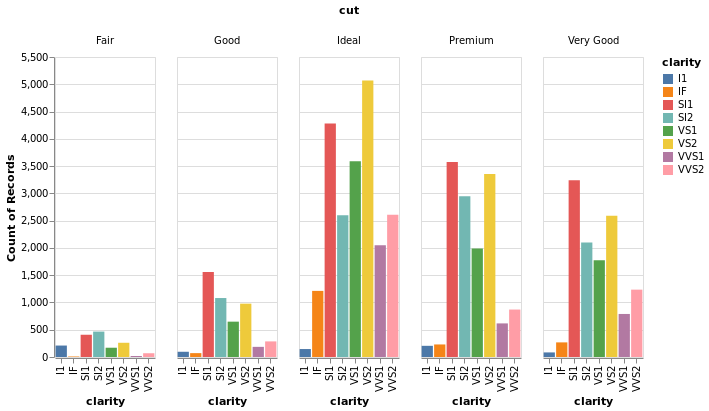
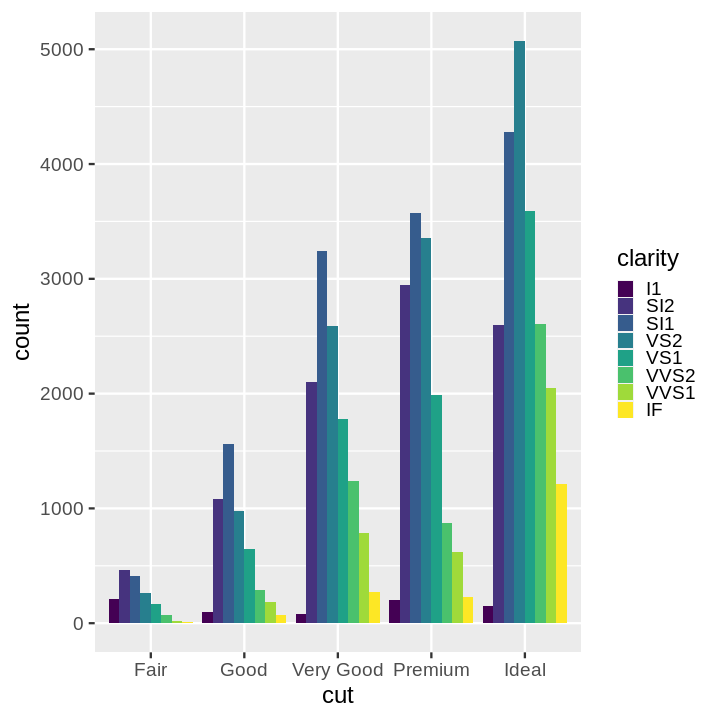
- Stacked Bar Chart
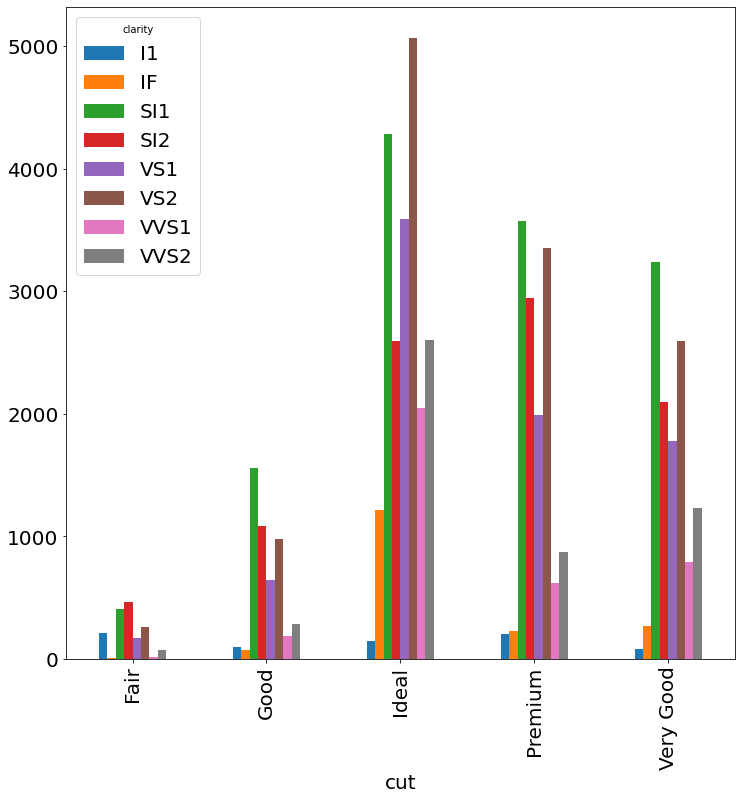
- Dodged Bar Chart
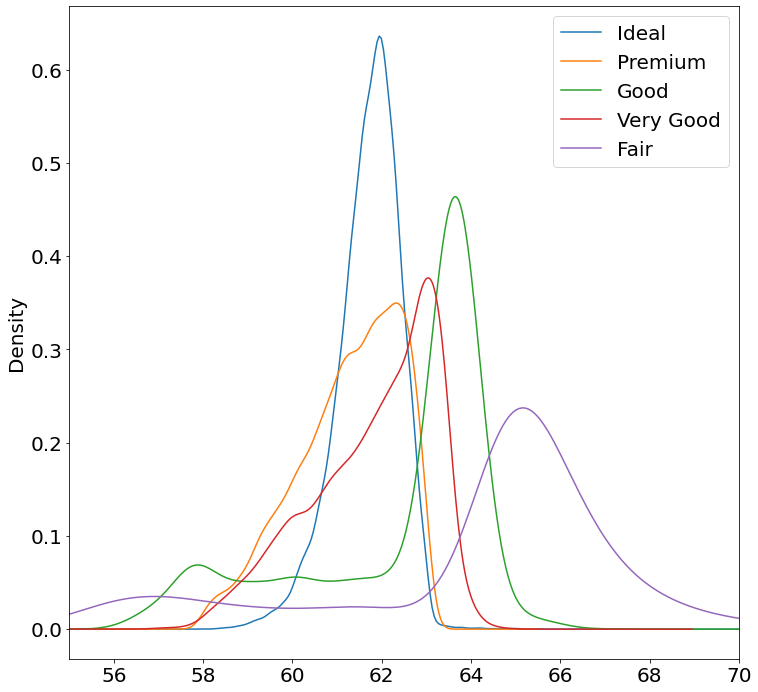
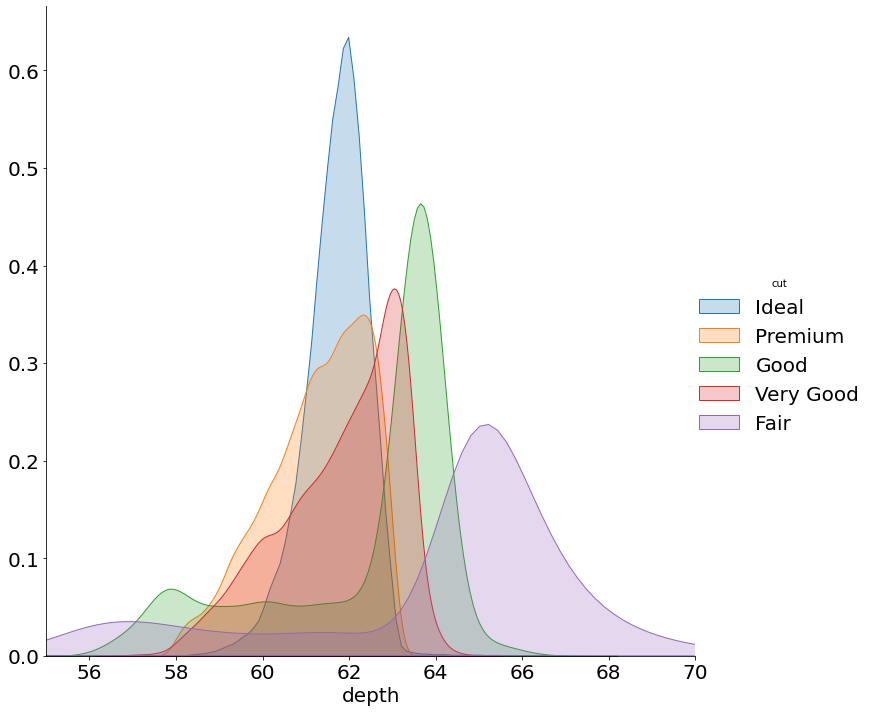
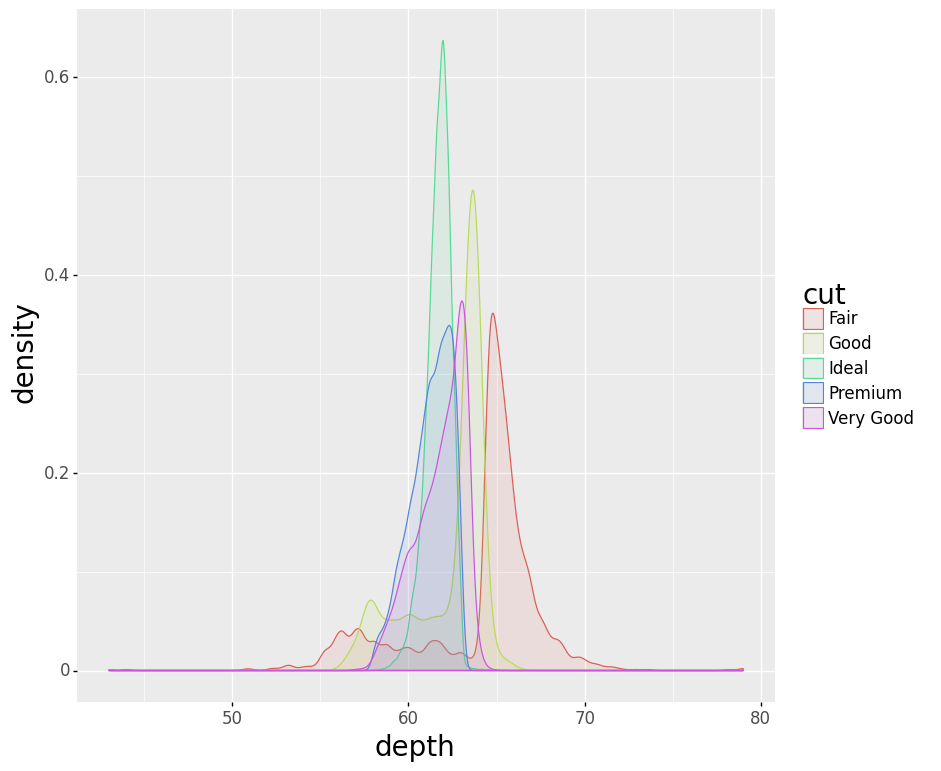
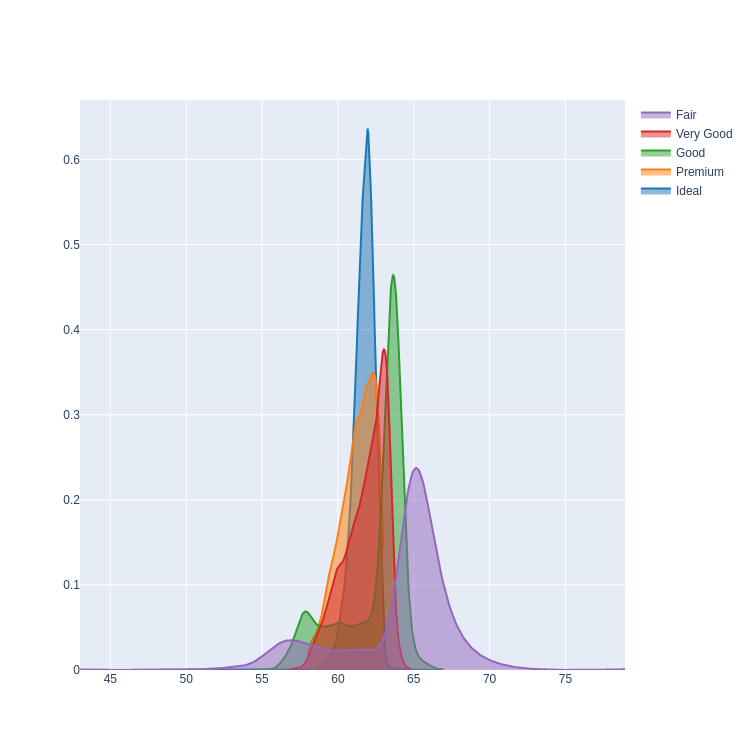
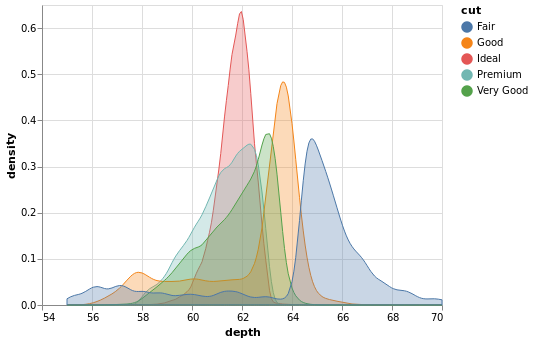
- Stacked KDE Plot
Introduction
Plotting is an essential component of data analysis. As a data scientist, I spend a significant amount of my time making simple plots to understand complex data sets (exploratory data analysis) and help others understand them (presentations).
In particular, I make a lot of bar charts (including histograms), line plots (including time series), scatter plots, and density plots from data in Pandas data frames. I often want to facet these on various categorical variables and layer them on a common grid.
Python Plotting Options
Python plotting libraries are manifold. Most well known is Matplotlib.
"Matplotlib is a Python 2D plotting library which produces publication quality figures in a variety of hardcopy formats and interactive environments across platforms." Native Matplotlib is the cause of frustration to many data analysts due to the complex syntax. Much of that frustration would be alleviated if it were recognized as a library of lower level plotting primitives on which other tools can be built. (If you are frustrated by Matplotlib and haven't read Effectively Using Matplotlib by Chris Moffitt, go read it.)
Matplotlib-Based Libraries
Many excellent plotting tools are built on top of Matplotlib.
Pandas plots provides the "basics to easily create decent looking plots" from data frames. It provides about 70% of what I want to do day-to-day. Importantly, it lacks robust faceting capabilities.
"plotnine is an implementation of a grammar of graphics in Python, it is based on ggplot2." plotnine is a attempt to directly translate ggplot2 to Python; despite some quirks and bugs, it works very well for a young product.
"Seaborn is a Python visualization library based on matplotlib. It provides a high-level interface for drawing attractive statistical graphics." Seaborn makes beautiful plots but is geared toward specific statistical plots, not general purpose plotting. It does have a powerful faceting utility function that I use regularly.
Interactive Plotting Libraries
There are several tools that can make the kinds of plots described here. At present, I have little experience with them. If anyone would like to help add examples, please get in touch.
"Altair is a declarative statistical visualization library for Python, based on Vega-Lite." According to Jake Vanderplas, "Declarative visualization lets you think about data and relationships, rather than incidental details." I provide Altair examples rendered as static images.
"plotly's Python graphing library makes interactive, publication-quality graphs online. Examples of how to make line plots, scatter plots, area charts, bar charts, error bars, box plots, histograms, heatmaps, subplots, multiple-axes, polar charts, and bubble charts." I provide plotly examples rendered as static images.
"Bokeh is a Python interactive visualization library that targets modern web browsers for presentation."
"bqplot is a Grammar of Graphics-based interactive plotting framework for the Jupyter notebook."
The Python Plotting Landscape
If you're interested in the breadth of plotting tools available for Python, I commend Jake Vanderplas's Pycon 2017 talk called the The Python Visualization Landscape. Similarly, the blogpost A Dramatic Tour through Python's Data Visualization Landscape (including ggplot and Altair) by Dan Saber is worth your time.
Hearty Thank You
Much Python plotting development is done by open source developers who have an (almost) thankless task. I am extremely grateful for the countless hours of many who have helped me do my job. Please keep it up!
Why all the talk about ggplot?
The word "ggplot" comes up a lot in discussions of plotting. Before I started using Python, I did most of my data analysis work in R. I, with many Pythonistas, remain a big fan of Hadley Wickham's ggplot2, a "grammar of graphics" implementation in R, for exploratory data analysis.
Like scikit-learn for machine learning in Python, ggplot2 provides a consistent API with sane defaults. The consistent interface makes it easier to iterate rapidly with low cognitive overhead. The sane defaults makes it easy to drop plots right into an email or presentation.
Particularly, ggplot2 allows the user to make basic plots (bar, histogram, line, scatter, density, violin) from data frames with faceting and layering by discrete values.
An excellent introduction to the power of ggplot2 is in Hadley Wickham and Garrett Grolemund's book R for Data Science.
Humble Rosetta Stone for Visualization in Exploratory Data Analysis
Below I have begun compiling a list of basic plots for exploratory data analysis. I have generated the plots with as many different libraries as time (and library) permits.
My hope is that this will (1) help you in your daily practice to work with what is available and (2) help inspire future development of Python plotting libraries.
Some rudimentary instructions on how you can contribute plots are here. General feedback or other plot suggestions are welcome.
Data
The datasets used below are included with ggplot2. One is the Prices of 50,000 round cut diamonds and the other is Fuel economy data from 1999 and 2008 for 38 popular models of car.
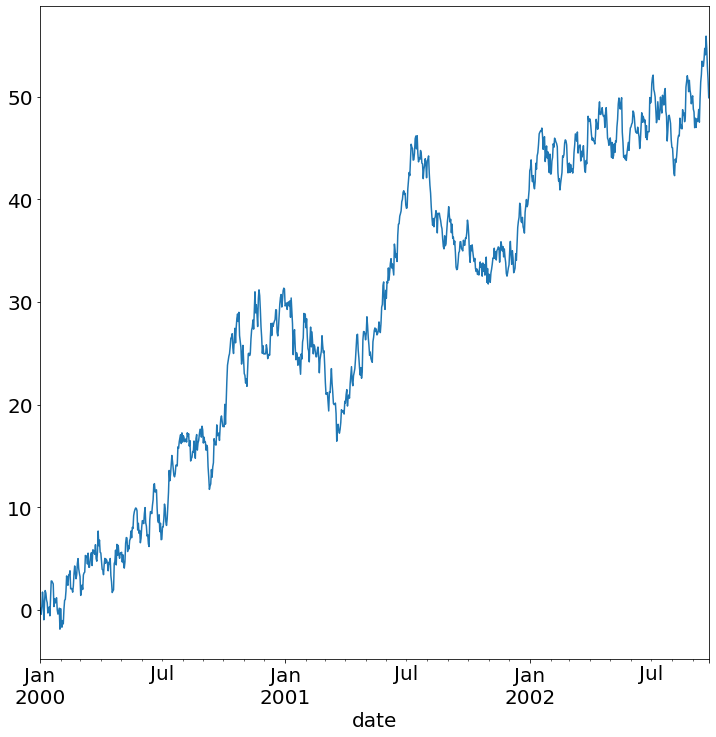
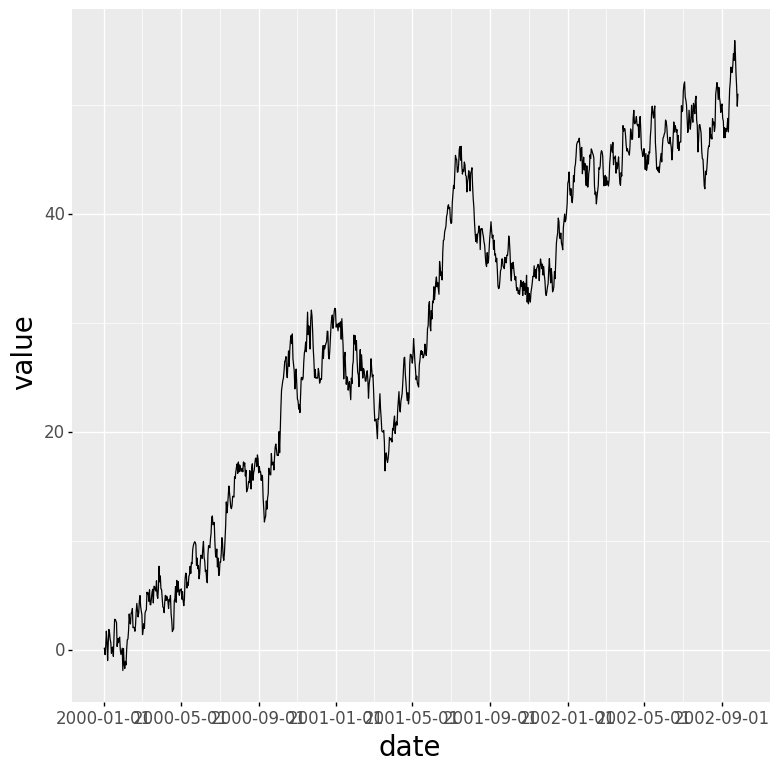
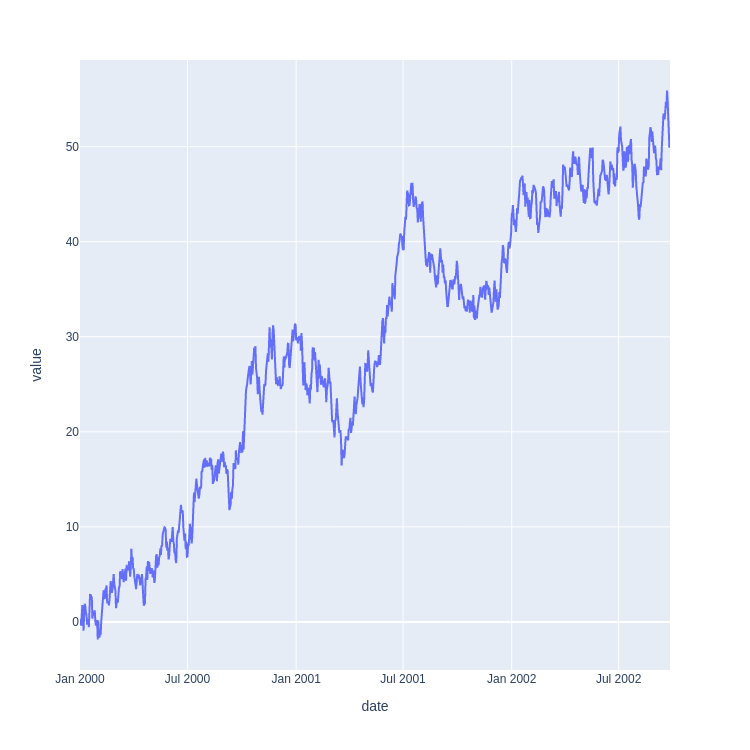
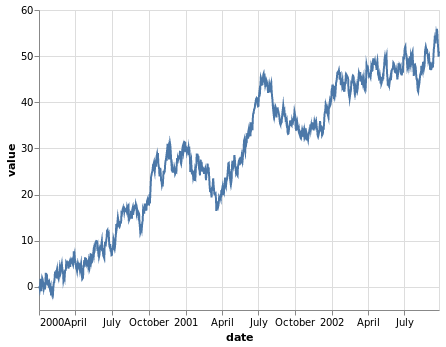
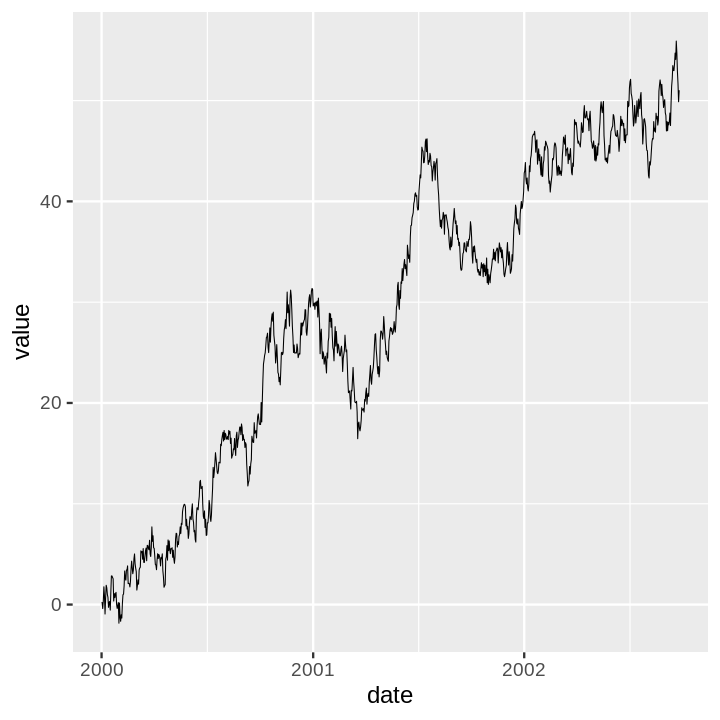
The time series example is a random walk I generate with a quick Python script.
Here's what a few rows of the datasets looks like:
ts
| date | value |
|---|---|
| 2000-01-01 | 0.218938 |
| 2000-01-02 | 0.195322 |
| 2000-01-03 | -0.397765 |
| 2000-01-04 | 0.359213 |
| 2000-01-05 | 1.760460 |
mpg
| manufacturer | model | displ | year | cyl | trans | drv | cty | hwy | fl | class |
|---|---|---|---|---|---|---|---|---|---|---|
| audi | a4 | 1.8 | 1999 | 4 | auto(l5) | f | 18 | 29 | p | compact |
| audi | a4 | 1.8 | 1999 | 4 | manual(m5) | f | 21 | 29 | p | compact |
| audi | a4 | 2.0 | 2008 | 4 | manual(m6) | f | 20 | 31 | p | compact |
| audi | a4 | 2.0 | 2008 | 4 | auto(av) | f | 21 | 30 | p | compact |
| audi | a4 | 2.8 | 1999 | 6 | auto(l5) | f | 16 | 26 | p | compact |
diamonds
| carat | cut | color | clarity | depth | table | price | x | y | z |
|---|---|---|---|---|---|---|---|---|---|
| 0.23 | Ideal | E | SI2 | 61.5 | 55.0 | 326 | 3.95 | 3.98 | 2.43 |
| 0.21 | Premium | E | SI1 | 59.8 | 61.0 | 326 | 3.89 | 3.84 | 2.31 |
| 0.23 | Good | E | VS1 | 56.9 | 65.0 | 327 | 4.05 | 4.07 | 2.31 |
| 0.29 | Premium | I | VS2 | 62.4 | 58.0 | 334 | 4.20 | 4.23 | 2.63 |
| 0.31 | Good | J | SI2 | 63.3 | 58.0 | 335 | 4.34 | 4.35 | 2.75 |
Code:
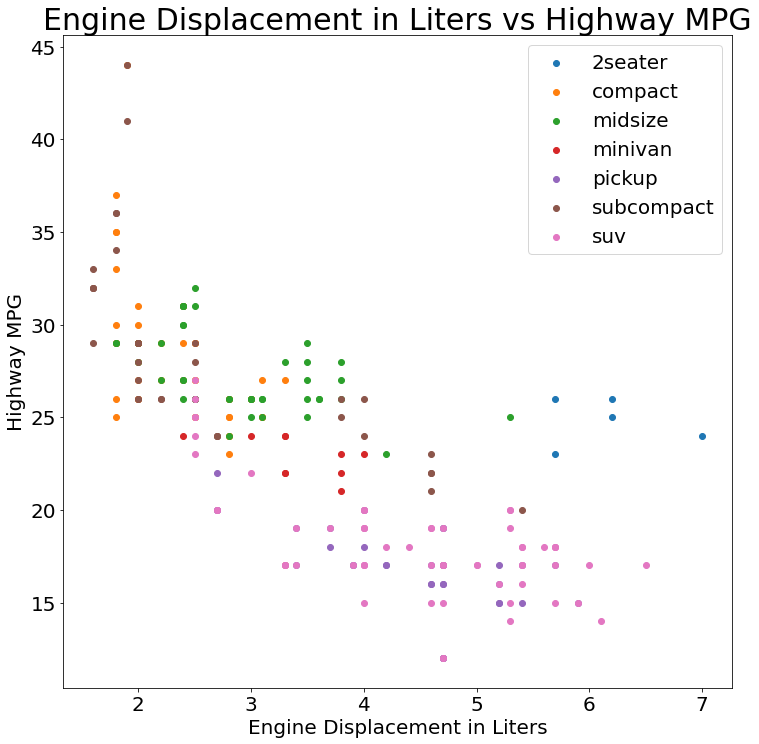
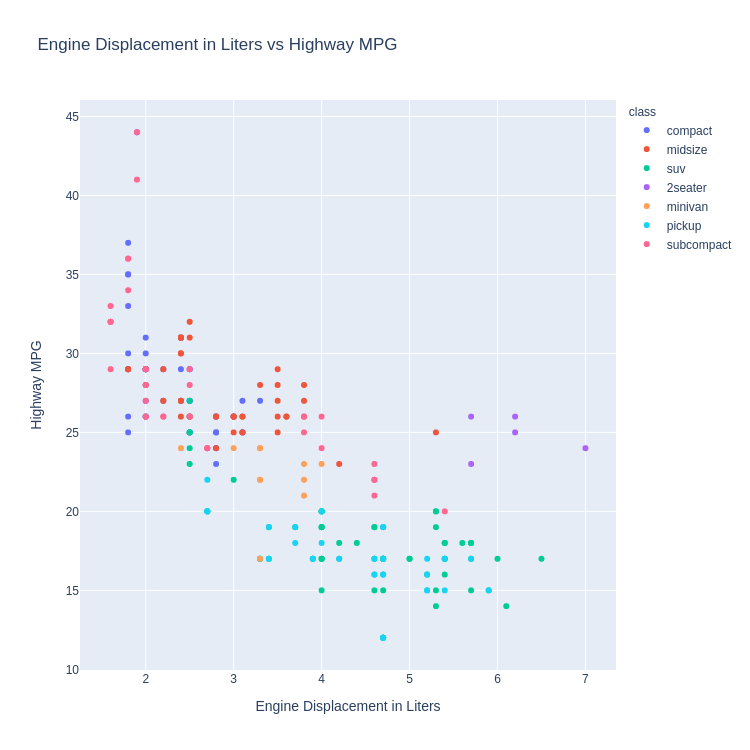
(sns
.FacetGrid(mpg, hue='class', size=10)
.map(pyplot.scatter, 'displ', 'hwy')
.add_legend()
.set(
title='Engine Displacement in Liters vs Highway MPG',
xlabel='Engine Displacement in Liters',
ylabel='Highway MPG'
))
Note:
seaborn.FacetGrid overrides the rcParams['figure.figsize'] global parameter.
You have to set the size in the size withsize=inFacetGrid`
Code:
import statsmodels.api as sm
from statsmodels.stats.outliers_influence import summary_table
y=mpg.hwy
x=mpg.displ
X = sm.add_constant(x)
res = sm.OLS(y, X).fit()
st, data, ss2 = summary_table(res, alpha=0.05)
preds = pd.DataFrame.from_records(data, columns=[s.replace('\n', ' ') for s in ss2])
preds['displ'] = mpg.displ
preds = preds.sort_values(by='displ')
fig = graph_objects.Figure(layout={
'title' : 'Engine Displacement in Liters vs Highway MPG',
'xaxis' : {
'title' : 'Engine Displacement in Liters'
},
'yaxis' : {
'title' : 'Highway MPG'
}
})
p1 = graph_objects.Scatter(**{
'mode' : 'markers',
'x' : mpg.displ,
'y' : mpg.hwy,
'name' : 'Points'
})
p2 = graph_objects.Scatter({
'mode' : 'lines',
'x' : preds['displ'],
'y' : preds['Predicted Value'],
'name' : 'Regression',
})
#Add a lower bound for the confidence interval, white
p3 = graph_objects.Scatter({
'mode' : 'lines',
'x' : preds['displ'],
'y' : preds['Mean ci 95% low'],
'name' : 'Lower 95% CI',
'showlegend' : False,
'line' : {
'color' : 'white'
}
})
# Upper bound for the confidence band, transparent but with fill
p4 = graph_objects.Scatter( {
'type' : 'scatter',
'mode' : 'lines',
'x' : preds['displ'],
'y' : preds['Mean ci 95% upp'],
'name' : '95% CI',
'fill' : 'tonexty',
'line' : {
'color' : 'white'
},
'fillcolor' : 'rgba(255, 127, 14, 0.3)'
})
fig.add_trace(p1)
fig.add_trace(p2)
fig.add_trace(p3)
fig.add_trace(p4)
Note:
No built in method to calculate and display confidence intervals. Must calculate manually and utilise existing features to build the confidence band.
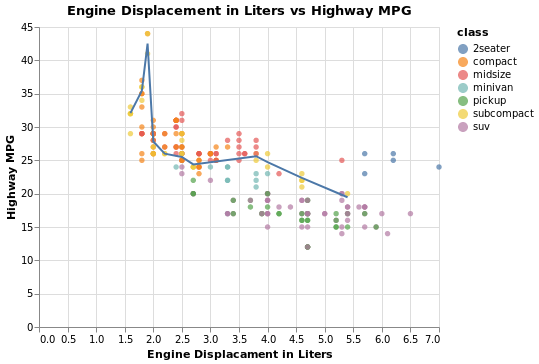
Code:
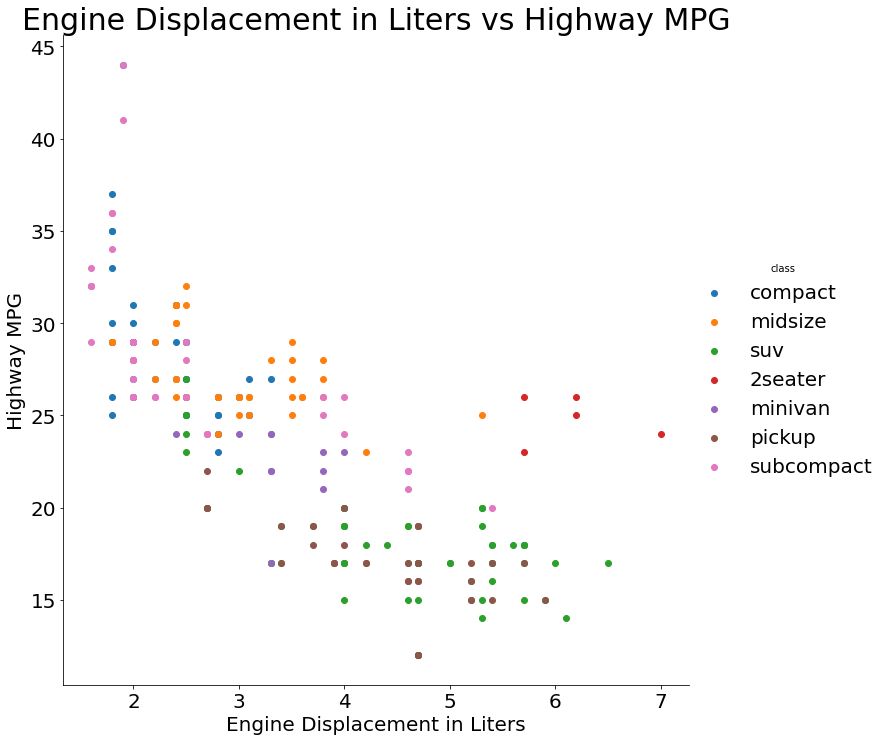
traces = []
for cls in mpg['class'].unique():
traces.append(graph_objects.Scatter({
'mode' : 'markers',
'x' : mpg.displ[mpg['class'] == cls],
'y' : mpg.hwy[mpg['class'] == cls],
'name' : cls
}))
subcompact = mpg[mpg['class'] == 'subcompact'].sort_values(by='displ')
traces.append(graph_objects.Scatter({
'mode' : 'lines',
'x' : subcompact.displ,
'y' : subcompact.hwy,
'name' : 'smoothing',
'line' : {
'shape' : 'spline',
'smoothing' : 1.3
}
}))
fig = graph_objects.Figure(**{
'data' : traces,
'layout' : {
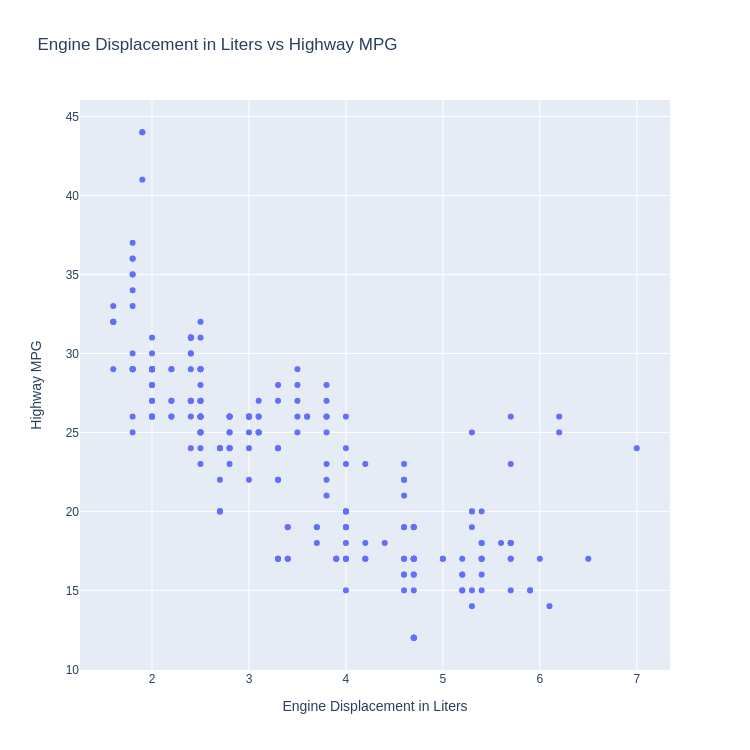
'title' : 'Engine Displacement in Liters vs Highway MPG',
'xaxis' : {
'title' : 'Engine Displacement in Liters',
},
'yaxis' : {
'title' : 'Highway MPG'
}
}
})
Note:
Plotly's builtin smoothing function is very weak
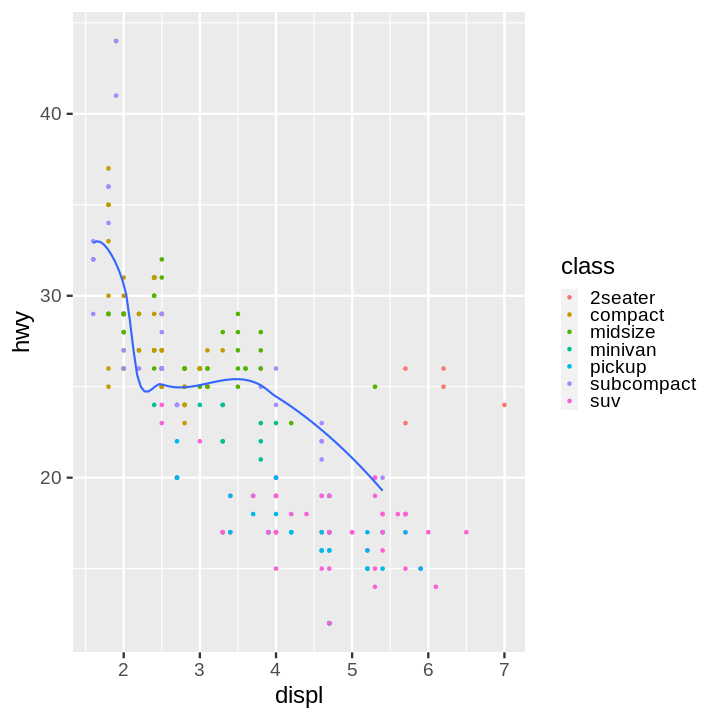
Code:
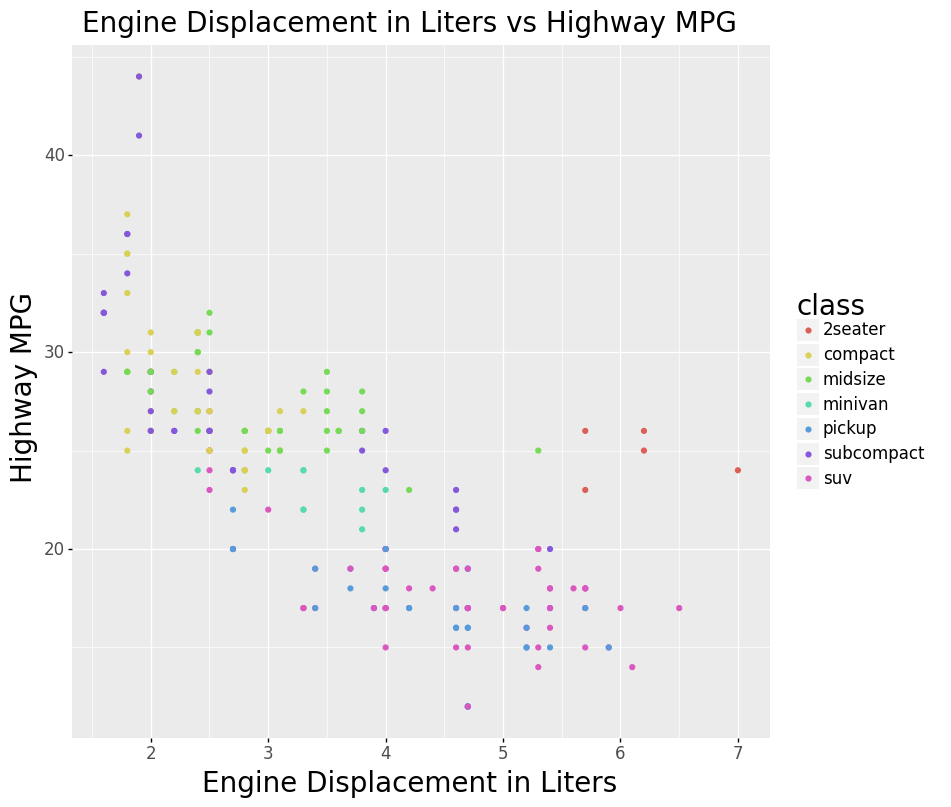
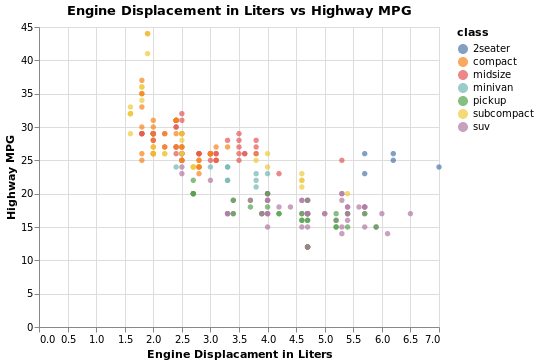
scatter = (
alt.Chart(
mpg,
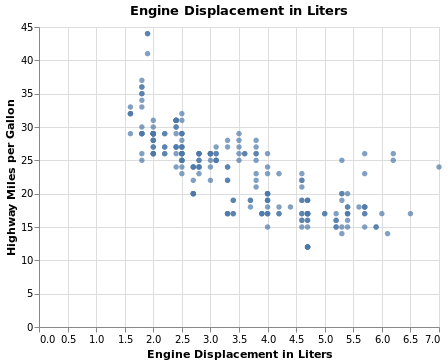
title='Engine Displacement in Liters vs Highway MPG',
)
.mark_circle()
.encode(
x=alt.X(
'displ',
axis=alt.Axis(
title='Engine Displacament in Liters'
),
),
y=alt.Y(
'hwy',
axis=alt.Axis(
title='Highway MPG'
),
),
color='class',
)
)
line = (
alt.Chart(
mpg[mpg['class'] == 'subcompact']
)
.transform_loess('displ', 'hwy')
.mark_line()
.encode(x=alt.X('displ'), y=alt.Y('hwy'))
)
scatter + line
Code:
fig, ax = pyplot.subplots()
ax.set_xlim(55, 70)
for cut in diamonds['cut'].unique():
s = diamonds[diamonds['cut'] == cut]['depth']
s.plot.kde(ax=ax, label=cut)
ax.legend()
Note:
I don't know whether Pandas can fill a KDE curve.
This requires using some Matplotlib to get them to stack and to have a legend.
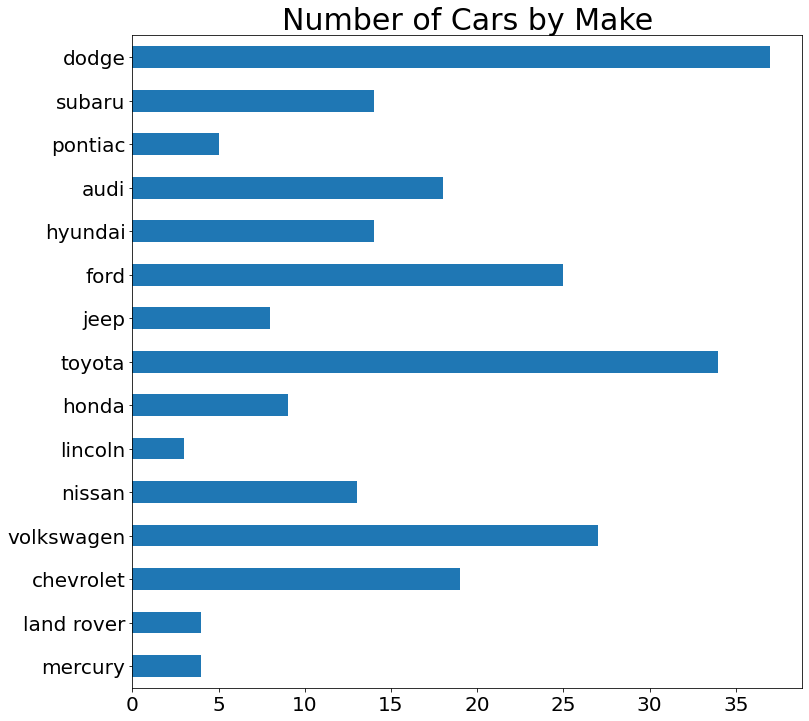
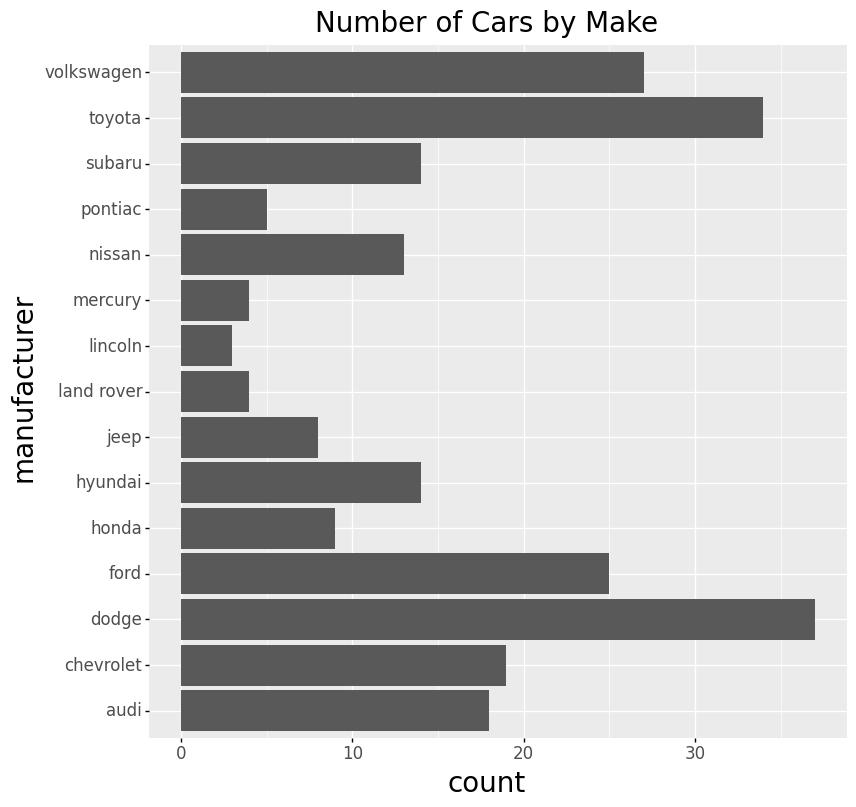
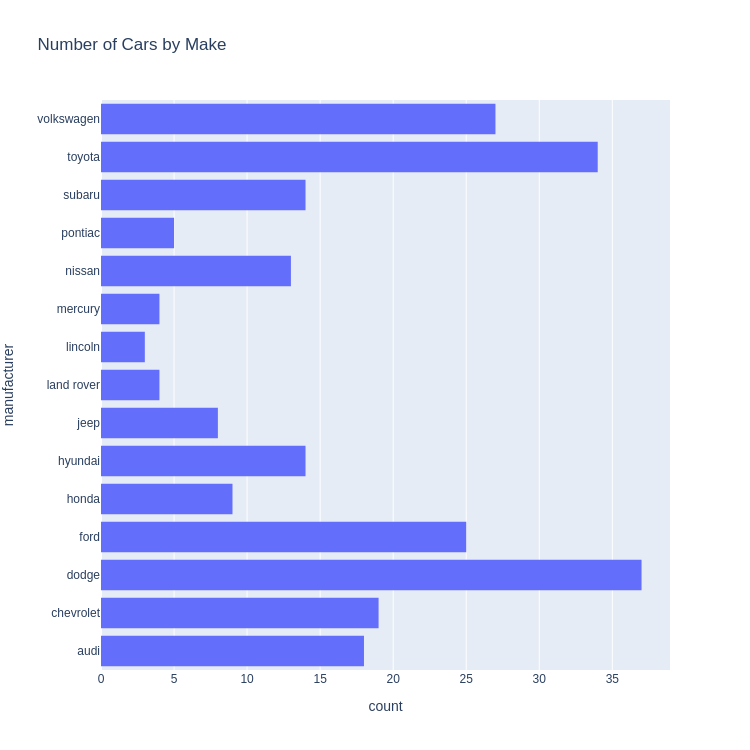
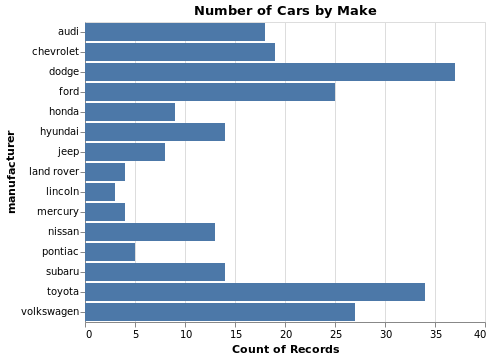
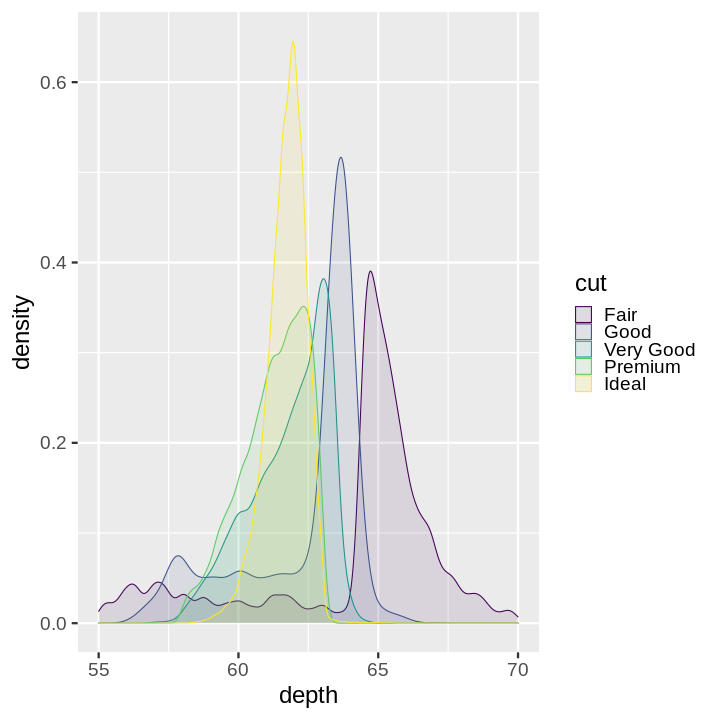
Note:
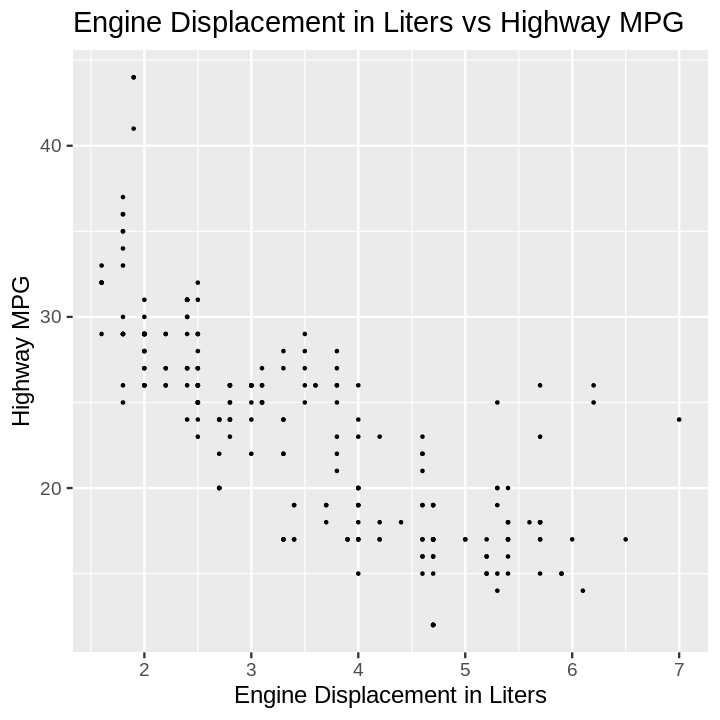
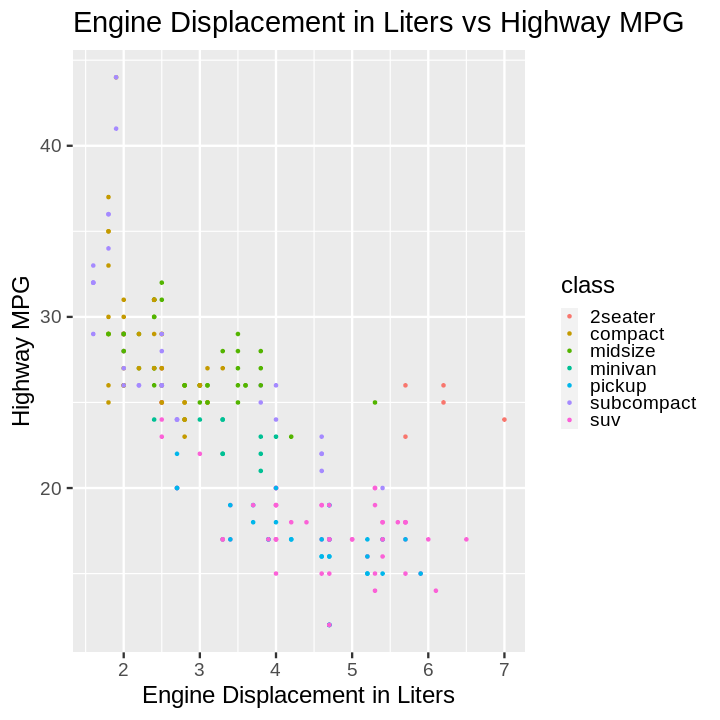
plotnine gives an error on
ggplot(data=mpg).